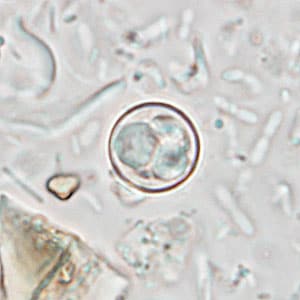
Oocyst of C. cayetanensis. In order to sequence these parasites, scientists needed new laboratory techniques that maximize DNA recovery from clinical samples because they cannot be grown in the lab. They also have a hard, resilient exterior that makes them difficult to break open to extract the DNA.
Updated on Tuesday, August 8, 2023
This discovery happened after nearly 10 years of improving laboratory methods, developing a special genotyping tool, and sequencing thousands of isolates.
In a recent study supported by AMD funding, researchers in CDC’s Division of Parasitic Diseases and Malaria (DPDM) describe that Cyclospora cayetanensis is not one singular organism, but at least three similar yet distinctive single-celled microscopic animals.
Cyclosporiasis is a debilitating intestinal illness caused when people eat food or drink water contaminated with a microscopic parasite called Cyclospora cayetanensis. In the United States, foodborne outbreaks of cyclosporiasis have been linked to various types of imported fresh produce, such as raspberries, basil, snow peas, mesclun lettuce, and cilantro. While some infected people have no symptoms at all, the vast majority suffer severe diarrhea and flu-like symptoms. Typically, the disease is treated with antibiotics, rest, and fluids, but, for some people, it can last a month or longer with frequent relapses.
Over the last 10 years, CDC scientists in DPDM developed laboratory techniques specifically to extract a sufficient amount of DNA for analysis from this tiny organism and created a new typing tool to compare the pathogen’s genome. Over the course of three years, 2018-2020, researchers genotyped thousands of isolates. They uncovered evidence that the three Cyclospora populations do not reproduce with one another, meaning that they are not reproductively compatible. Without the genotyping data, these patterns—and their potential benefit for public health—would have remained undetected.
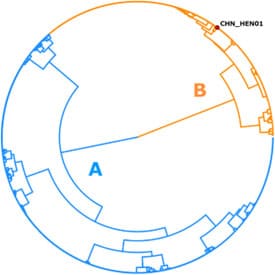
Dr. Joel Barratt, a bioinformatician and molecular parasitologist with DPDM, says this discovery allows scientists and public health workers to discuss the organisms by unique names, much as it is possible to discuss and differentiate strains of SARS-CoV-2. As researchers studied the differences, they also found evidence that each of the species tends to cause infection in specific parts of the world.
In the United States, preliminary data seems to indicate that one is more prevalent in southern states, while another causes infection more frequently in upper midwestern states. The third species has only been detected in China. Moreover, in the US, infections seem to happen at slightly different times of year depending on which species is making people sick. This might be related to supply chains for fresh produce. Continued analysis will help public health officials trace the source of an outbreak based on the distinctive genetic description of the Cyclospora species involved and could determine which produce items are more likely to be associated with a particular species. With surveillance, scientists can also determine if a particular species passes from person to person more easily or makes people sicker.
DPDM scientists have been using AMD technologies to better study the complex genetics of Cyclospora with support from the AMD program since the program launched in 2014. Now, CDC’s C. cayetanensis genotyping system is used to complement outbreak investigations and has a growing library of data. “The funding is significant not only in the development of routine public health programs like the Cyclospora genotyping program, but also in facilitating the generation of new scientific knowledge on pathogens of public health interest,” says Dr. Barratt.
Lineage: Cyclospora cayetanensis (aka Lineage A)
Locality: Cyclospora cayetanensis causes infections in the Americas, though may infect humans elsewhere.
Lineage: Cyclospora ashfordi (aka Lineage B)
Locality: Cyclospora ashfordi causes infections in the Americas, though may infect humans elsewhere.
Lineage: Cyclospora henanensis (aka Lineage C)
Locality: The strain caused an infection in Henan province, China, though may infect humans elsewhere.