Module 2.1 – SARS-CoV-2 sequencing in Arizona
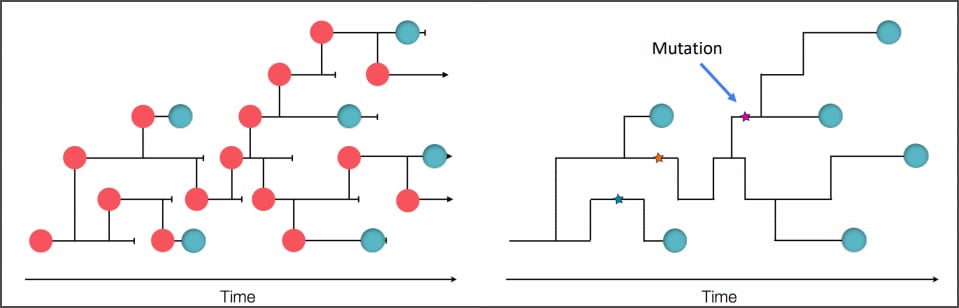
Overview: This module provides insight into how SARS-CoV-2 sequencing is used to describe the genomic epidemiology of a state and as an investigative tool in COVID-19 outbreak settings. You can read more about this work: An Early Pandemic Analysis of SARS-CoV-2 Population Structure and Dynamics in Arizonaexternal icon
Posted: 01/08/21
Presenter: Hayley Yaglom (view bio), MS, MPH
Genomic Epidemiologist, Translational Genomics Research Institute
View Presentation [Full Version]external icon [Short Version]external icon
Further Reading:
- An early pandemic analysis of SARS-CoV-2 population structure and dynamics in Arizona.external icon
Ladner, et al. mBio, 2020. - AZ-Strain: Genomic epidemiology of SARS-CoV-2 in Arizona.external icon
Arizona COVID Genomics Union (ACGU).
Additional Resources:
- Communicating results using narratives.external icon
Nextstrain.org (documentation). - Coast-to-coast spread of SARS-CoV-2 during the early epidemic in the United States.external icon
Fauver, et al. Cell, 2020. - Large-scale sequencing of SARS-CoV-2 genomes from one region allows detailed epidemiology and enables local outbreak management.external icon
Page, et al. Microb Genom, 2021. - Viral genomes reveal patterns of the SARS-CoV-2 outbreak in Washington State.external icon
Mueller, et al. Sci Transl Med, 2021. - Revealing fine-scale spatiotemporal differences in SARS-CoV-2 introduction and spread.external icon
Moreno, et al. Nat Commun, 2020.
Take the Feedback Survey for Module 2.1
email_03Get Email Updates
More modules and materials will be added to this toolkit, so please check back for updates or subscribe to the mailing list.
Related Videos
Page last reviewed: September 3, 2021
Content source: Centers for Disease Control and Prevention