FoodCORE Year Eight Summary
January 1, 2018 to December 31, 2018
Background
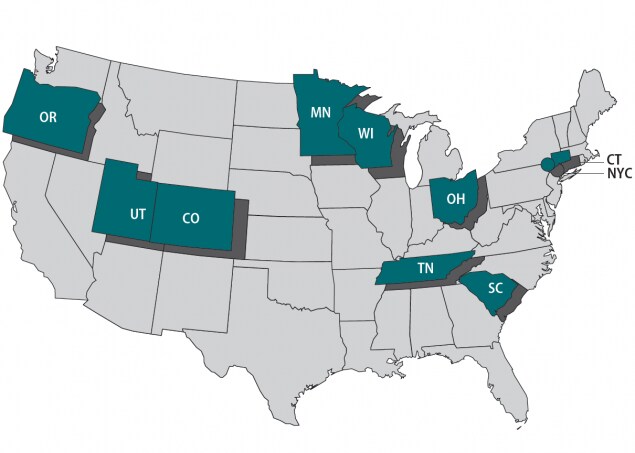
Foodborne Diseases Centers for Outbreak Response Enhancement (FoodCORE) centers address gaps in foodborne disease response through enhanced capacity in laboratory, epidemiology, and environmental health to improve timeliness and completeness of outbreak response activities. The FoodCORE centers during Year Eight (January 1, 2018 – December 31, 2018) were Colorado, Connecticut, Minnesota, New York City, Ohio, Oregon, South Carolina, Tennessee, Utah, and Wisconsin.
Program Highlights
In Year Eight, the FoodCORE centers revised the Initial Case-patient Interviewing Model Practice. This model practice, originally posted to the FoodCORE website in 2014, describes the methods used by FoodCORE centers to gather detailed exposure histories from case-patients. Since the document was first written, new investigation methods have emerged. The 2018 version of this model practice expanded the section on reaching case-patients to include texting and the use of online questionnaires. These newer methods and the strategies described in the original document have allowed FoodCORE centers to attempt interviews with nearly all case-patients and reduce the proportion of cases that are lost to follow up.

While their efforts primarily focus on foodborne outbreaks, FoodCORE centers’ enhanced capacity has successfully aided in the response to outbreaks related to other sources and etiologies. One success story highlighted two zoonotic outbreaks in FoodCORE centers – a turtle-associated outbreak in Ohio and a goat-associated outbreak in Connecticut. Another success story highlighted Colorado, New York City, Ohio, and Utah all supporting non-foodborne outbreaks. They investigated outbreaks spread by animals, sexual contact, and throughout a community that included individuals experiencing homelessness.
In January 2018, whole genome sequencing (WGS) became the standard subtyping method supported by PulseNet for Listeria isolates. FoodCORE centers prepared for the transition and sequenced 100% of their Listeria isolates in 2018. In 2019, WGS will become the standard subtyping method for Escherichia coli, Shigella, and Salmonella. FoodCORE metrics will measure this transition and the impact of WGS on timeliness and completeness of subtyping as well as cluster and outbreak detection.
To share all of these great successes, FoodCORE staff at CDC and FoodCORE centers presented at several meetings and conferences, including the International Conference on Emerging Infectious Diseases and the Council for State and Territorial Epidemiologists Annual Conference.
Program Performance
Centers report metrics twice a year to document changes resulting from targeted FoodCORE resources. Metrics for Salmonella, Shiga toxin-producing Escherichia coli (STEC), and Listeria (SSL) have been collected since late 2010. Metrics for norovirus, other etiologies, and unknown etiology (NOU) investigations have been collected since 2012. The metrics collected by FoodCORE centers are revised as needed to best meet program goals. See below for figures and graphs for selected metrics. Information on all of the metrics and complete data tables are available on the FoodCORE website.
Graphs for Selected Metrics – Year Eight
Since Year 6, FoodCORE centers have increased the proportion of Salmonella, STEC, and Listeria primary isolates with WGS results.
To evaluate the timeliness and completeness of WGS, FoodCORE centers pilot-tested a set of expanded SSL metrics in Year 6. Centers reported on the expanded metrics in Year 6 and in following years. Download and view chart data excel icon[XLS – 38.5 KB].
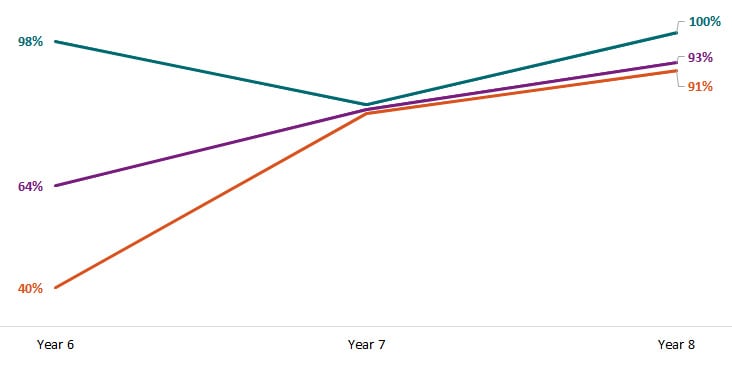
Orange = Salmonella. Purple = STEC. Teal = Listeria.
From Year 7 to Year 8, FoodCORE centers reduced the time* from Salmonella, STEC, and Listeria isolate receipt (or recovery) at the public health lab to WGS sequence being shared with the national database.
Download and view chart data excel icon[XLS – 38.5 KB].
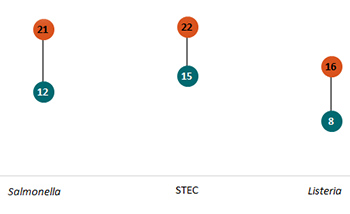
Orange = Year 7. Teal = Year 8. *Time in median days.
FoodCORE centers contacted a greater proportion of environmental health, agriculture, regulatory, consumer protection, or food safety program staff for foodborne or point-source investigations in Year 8 than in previous years.
Download and view chart data excel icon[XLS – 38.5 KB].
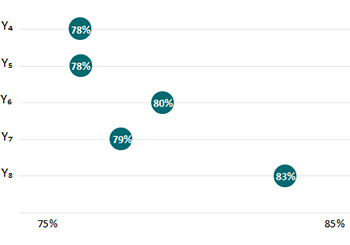
Year 4 (Y4) = January 1, 2014–December 31, 2014. Year 5 (Y5) = January 1, 2015–December 31, 2015. Year 6 (Y6) = January 1, 2016–December 31, 2016. Year 7 (Y7) = January 1, 2017–December 31, 2017. Year 8 (Y8) = January 1, 2018–December 31, 2018.
In Year 8, the time* to attempt and complete interviews for Salmonella, STEC, and Listeria was under 2 days and 4 days, respectively.
Download and view chart data excel icon[XLS – 38.5 KB].
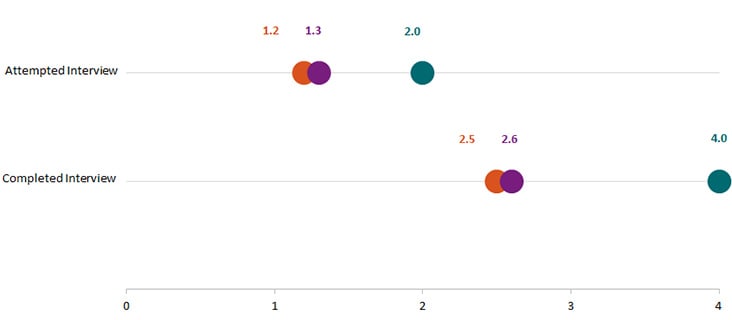
Orange = Salmonella. Purple = STEC. Teal = Listeria. *Time in median days.
Center-specific Accomplishments and Investigations in Year 8
| FoodCORE Reporting Periods | |||
| Baseline (Y0) =
Oct 2010 – Mar 2011 |
Year 1 (Y1) =
Oct 2010 – Sept 2011 |
Year 2 (Y2) =
Oct 2011 – Dec 2012 |
Year 3 (Y3) =
Jan 2013 – Dec 2013 |
| Year 4 (Y4) =
Jan 2014 – Dec 2014 |
Year 5 (Y5) =
Jan 2015 – Dec 2015 |
Year 6 (Y6) =
Jan 2016 – Dec 2016 |
Year 7 (Y7) =
Jan 2017 – Dec 2017 |
| Year 8 (Y8) =
Jan 2018 – Dec 2018 |
|||
FoodCORE centers have demonstrated that targeted investments can improve the completeness and timeliness of outbreak response activities. They have strengthened their outbreak response programs to conduct faster, better, and more complete investigations, to ultimately help stop the spread of infectious diseases. Download a print version of the FoodCORE Year Eight Summary pdf icon[PDF – 683 KB].

Download a print version of the FoodCORE Year Eight Summary pdf icon[PDF - 683 KB].