Influenza Virus Genome Sequencing and Genetic Characterization
Genome Sequencing
Influenza viruses are constantly evolving, in fact all influenza viruses undergo genetic changes over time (for more information, see How the Flu Virus Can Change: “Drift” and “Shift”). An influenza virus’ genome consists of all genes that make up the virus. CDC conducts year-round surveillance of circulating influenza viruses to monitor changes to the genome (or parts of the genome) of these viruses. This work is performed as part of routine U.S. influenza surveillance and as part of CDC’s role as a World Health Organization (WHO) Collaborating Center for Reference and Research on Influenza. The information CDC collects from studying genetic changes (also known as “substitutions,” “variants” or “mutations”) in influenza viruses plays an important public health role by helping to determine whether existing vaccines and medical countermeasures (e.g., antiviral drugs) will work against new influenza viruses, as well as helping to determine the potential for influenza viruses in animals to infect humans.
Genome sequencing reveals the sequence of the nucleotides in a gene, like alphabet letters in words. Comparing the composition of nucleotides in one virus gene with the order of nucleotides in a different virus gene can reveal variations between the two viruses.
Genetic variations are important because they affect the structure of an influenza virus’ surface proteins. Proteins are made of sequences of amino acids.
The substitution of one amino acid for another can affect properties of a virus, such as how well a virus transmits between people, and how susceptible the virus is to antiviral drugs or current vaccines.
Influenza A and B viruses – the primary influenza viruses that infect people – are RNA viruses that have eight gene segments. These genes contain ‘instructions’ for making new viruses, and it’s these instructions that an influenza virus uses once it infects a human cell to trick the cell into producing more influenza viruses, thereby spreading infection.
Influenza genes consist of a sequence of molecules called nucleotides that bond together in a chain-like shape. Nucleotides are designated by the letters A, U, C or G.
Genome sequencing is a process that determines the order, or sequence, of the nucleotides (i.e., A, U, C or G) in each of the genes present in the virus’s genome. Full genome sequencing can reveal the approximately 13,500-letter sequence of all the genes of the virus’ genome, while partial-genome sequencing reveals the sequence of some or parts of those genes.
Each year CDC performs whole genome sequencing on about 6,000 influenza viruses from original clinical samples collected through virologic surveillance. Of the eight genes that make up an influenza A or B virus, CDC focuses on sequencing two gene segments: the hemagglutinin (HA) and the neuraminidase (NA). The HA/NA genes determine the structure of the two primary surface proteins of influenza viruses. Surface proteins determine important properties of influenza viruses, including how the influenza virus responds to antiviral drugs, the virus’ genetic similarity to influenza vaccine viruses, and the potential for zoonotic (animal origin) influenza viruses to infect human hosts.
Genetic Characterization
CDC and other public health laboratories around the world have been sequencing the genes of influenza viruses since the 1980s. Today, such gene sequences are compiled in databases for use by public health researchers, such as GenBank and the Global Initiative on Sharing Avian Influenza Data (GISAID). The resulting libraries of gene sequences allow CDC and other laboratories to compare the genes of currently circulating influenza viruses with the genes of older influenza viruses and viruses used in vaccines. Through this process of comparing genetic sequences, called genetic characterization, CDC can make informed assumptions regarding:
- How flu viruses are ‘related’ to one another
- How flu viruses are evolving
- The genetic variations (a.k.a., substitutions or mutations) that appear when viruses begin spreading more easily, causing more-severe disease, or developing resistance to antiviral drugs
- How well an influenza flu vaccine might protect against a particular influenza virus
- Adaptations in influenza viruses circulating in animal populations that may enable the virus to infect humans.
The relative differences among a group of influenza viruses are shown by organizing them into a graphic called a ‘phylogenetic tree.’ Phylogenetic trees for influenza viruses are like family (genealogy) trees for people. These trees show how closely ‘related’ individual viruses are to one another. Viruses are grouped together based on whether their genes’ nucleotides are identical or not. Phylogenetic trees of influenza viruses will usually display how similar the viruses’ hemagglutinin (HA) or neuraminidase (NA) genes are to one another. Each sequence from a specific influenza virus has its own branch on the tree. The degree of genetic difference (number of nucleotide differences) between viruses is represented by the length of the horizontal lines (branches) in the phylogenetic tree. The further apart viruses are on the horizontal axis of a phylogenetic tree, the more genetically different the viruses are to one another.
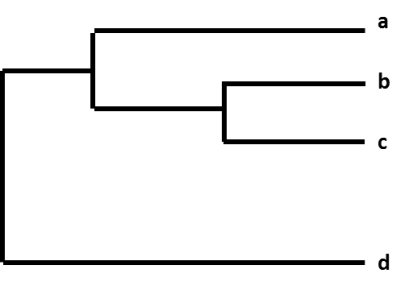
Figure. A phylogenetic tree.
For example, after CDC sequences an influenza A(H3N2) virus collected through surveillance, the virus sequence is cataloged with other virus sequences that have a similar HA gene (H3), and a similar NA gene (N2). As part of this process, CDC compares the new virus sequence with the other virus sequences, and looks for differences among them. CDC then uses a phylogenetic tree to visually represent how genetically different the H3N2 viruses are from each other.
CDC performs genetic characterization of influenza viruses year round. This genetic data is used in conjunction with virus antigenic characterization data to help determine which vaccine viruses should be chosen for the upcoming Northern Hemisphere or Southern Hemisphere influenza vaccines. In the months leading up to the WHO vaccine consultation meetings in February and September, CDC collects influenza viruses through surveillance and compares the HA and NA gene sequences of current vaccine viruses against those of circulating flu viruses. This is one way to assess how closely related the circulating influenza viruses are to the viruses the seasonal flu vaccine was formulated to protect against. As viruses are collected and genetically characterized, differences can be revealed.
For example, sometimes over the course of a season, circulating viruses will change genetically, which causes them to become distinctly different from the corresponding vaccine virus. This is one indication that a different vaccine virus may need to be selected for the next flu season’s vaccine, although other factors, including antigenic characterization findings, heavily influence vaccine decisions. The HA and NA surface proteins of influenza viruses are antigens, which means they are recognized by the immune system and are capable of triggering an immune response, including production of antibodies that can block infection. Antigenic characterization refers to the analysis of a virus’s reaction with antibodies to help assess how it relates to another virus.
Methods of Flu Genome Sequencing
One influenza sample contains many influenza virus particles that were grown in a test tube and that often have small genetic differences in comparison to one another among the whole population of sibling viruses.
Traditionally, scientists have used a sequencing technique called “the Sanger reaction” to monitor influenza evolution as part of virologic surveillance. Sanger sequencing identifies the predominant genetic sequence among the many influenza viruses found in an isolate. This means small variations in the population of viruses present in a sample are not reflected in the final result. Scientists often use the Sanger method to conduct partial genome sequencing of influenza viruses, while newer technologies (see next paragraph) are better suited for whole genome sequencing.
Over the past five years, CDC has been using “Next Generation Sequencing (NGS)” methodologies, which have greatly expanded the amount of information and detail that sequencing analysis can provide. Unlike Sanger sequencing, NGS uses advanced molecular detection (AMD) to identify gene sequences from each virus in a sample. Therefore, NGS reveals the genetic variations among many different influenza virus particles in a single sample, and these methods also reveal the entire coding region of the genomes. This level of detail can directly benefit public health decision-making in important ways, but data must be carefully interpreted by highly-trained experts in the context of other available information. See AMD Projects: Improving Influenza Vaccines for more information about how NGS and AMD are revolutionizing flu genome mapping at CDC.



