FoodNet Fast: Diagnostic Laboratory Practices Tool FAQ
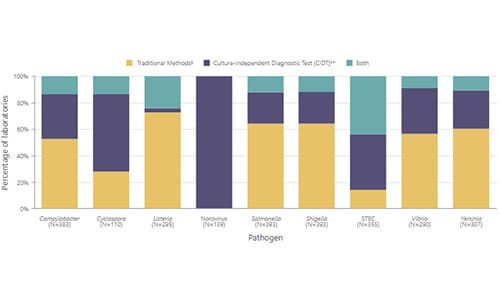
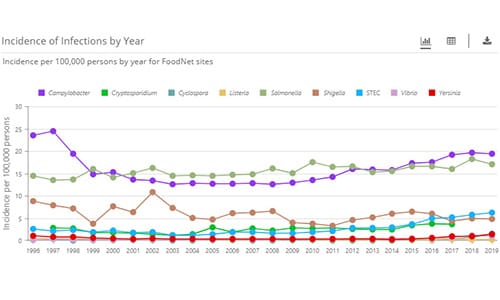
FoodNet Fast’s Diagnostic Laboratory Practices Tool lets you find out how testing practices of about 700 diagnostic laboratories in FoodNet’s surveillance area have changed over time for 10 pathogens transmitted commonly through food: Campylobacter, Cryptosporidium*, Cyclospora, Listeria, norovirus, Salmonella, Shiga toxin-producing E. coli (STEC), Shigella, Vibrio, and Yersinia.
*As of January 1, 2018, FoodNet no longer collects information on laboratory practices for Cryptosporidium.