Questions and Answers about Lab Testing

Questions in This Section
- What specimens should be collected from patients who meet the mumps clinical case definition?
- When is the optimal timing for collection of specimens for laboratory confirmation of mumps?
- What tests are available for laboratory confirmation of mumps?
- Where do I send laboratory specimens for testing?
- We have seen cases of “recurrent parotitis” in our state. Is this unusual and is it necessary to recommend isolation again from their last onset date?
Q: What specimens should be collected from patients who meet the mumps clinical case definition?
A: CDC recommends collection of a buccal swab specimen as soon after parotitis onset as possible from all patients with clinical features compatible with mumps. See Specimen Collection, Storage, and Shipment for detailed instructions regarding recommended specimen for mumps testing.
Q: When is the optimal timing for collection of specimens for laboratory confirmation of mumps?
A: A buccal swab specimen for detection of virus should be collected as soon as possible upon suspicion of mumps disease. The early collection of buccal swab specimens provides the best means of laboratory confirmation, particularly among suspected mumps patients with a history of vaccination (Rota et al, 2013). If it has been <3 days since symptom onset, collect only a buccal swab specimen for detection of viral RNA by rRT-PCR. If it has been >3 days since symptom onset, collect a buccal swab specimen for rRT- PCR and a serum specimen for IgM detection. If the patient has orchitis/oophoritis, mastitis, pancreatitis, hearing loss, meningitis, or encephalitis, collect a buccal swab specimen for rRT-PCR and a serum specimen for IgM detection, regardless of days since symptom onset.
In an outbreak setting (see Figure below), the proportion of specimen with positive results by rRT-PCR (N target gene) decreased as the interval between onset and specimen collection increased (Rota et al., 2013). The trend was reversed for IgM detection with increasing time to specimen collection the number of seropositive specimens increased.
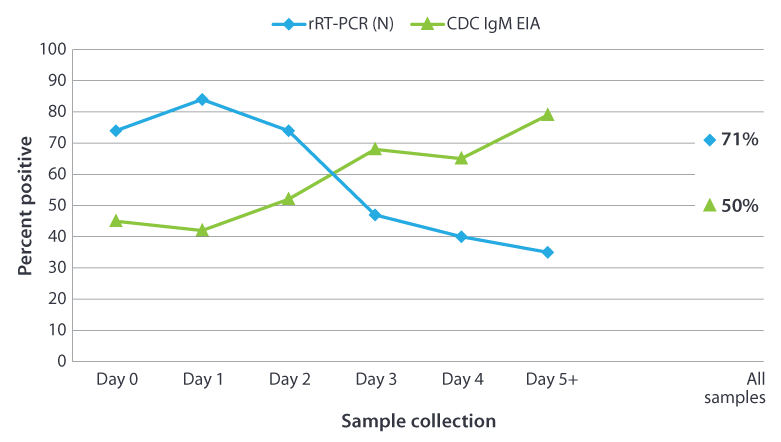
The percentage of positive results obtained from testing 296 confirmed mumps cases from New York City by day of sample collection after onset of symptoms. The serum specimen were tested for presence of IgM using the CDC capture IgM EIA. The buccal swab specimen were tested by rRT-PCR using the mumps nucleoprotein (N) gene as the target.
*Done in collaboration with New York City Department of Health and Mental Hygiene Public Health Laboratory, New York, NY
Mumps virus was isolated from 209 (71%) of the 296 buccal swabs tested.
Q: What tests are available for laboratory confirmation of mumps?
A: Viral detection methods include molecular assays, such as real time reverse transcription PCR (rRT-PCR) to detect mumps viral RNA. rRT-PCR is performed by almost all state and local public health laboratories, the APHL Vaccine Preventable Disease Reference Centers (VPD-RC [2 pages]), and CDC. Genotyping, based on the sequence of the gene coding for the small hydrophobic (SH) protein can also be performed by the VPD-RCs and CDC and this is the only test to discriminate vaccine reactions from wild-type infections.
IgM can aid in diagnosis but is not confirmatory. The availability of assays to detect IgM to aid in the diagnosis of acute mumps infection and to measure IgG antibodies to document previous exposure to mumps vary among laboratories. The state health department can provide guidance regarding available laboratory services. At the direction of the state health department, healthcare providers and state and local health departments may send serum specimens from suspected mumps cases to CDC/VVPDB for IgM detection using a capture IgM enzyme immunoassay that incorporates a recombinant mumps nucleocapsid protein as the antigen.
Q: Where do I send laboratory specimens for testing?
Q: We have seen cases of “recurrent parotitis” in our state. Is this unusual and is it necessary to recommend isolation again from their last onset date?
Detection of Mumps Virus in Clinical Specimen
Questions in This Section
- Is collection of both a buccal swab and a serum sample necessary?
- What does a mumps-negative rRT-PCR result mean?
- What does a mumps-positive rRT-PCR result mean?
- Why doesn’t a negative rRT-PCR result and/or a negative IgM result rule out mumps as a diagnosis?
- What is a mumps genotype?
- What mumps virus genotype has been commonly detected in the United States?
- Are there small genetic changes that may be affecting our molecular diagnostics?
- Would genetic changes that are occurring cause the vaccine not to work against certain strains of the virus?
- If a clinical sample is determined to be positive by rRT-PCR or by viral isolation at the state laboratory, is it necessary to send a sample to CDC for sequence analysis?
- Where can I find RT-PCR and real-time RT-PCR protocols for mumps?
- Where do I obtain material for rRT-PCR and virus isolation controls?
Q: Is collection of both a buccal swab and a serum sample necessary?
A: CDC recommends collection of a buccal swab specimen for rRT-PCR if it has been <3 days since symptom onset. Buccal/oral swab specimen enhance the ability to laboratory-confirm a mumps infection and can provide additional information (sequence analysis to obtain the mumps genotype) to aid epidemiologic investigations. In addition, because serologic tests cannot differentiate between an exposure to vaccine and an exposure to wild-type mumps virus, it is necessary to obtain buccal specimen to differentiate between wild-type and vaccine virus. If it has been >3 days since symptom onset, collect a buccal swab specimen for rRT- PCR and a serum specimen for IgM detection. This may be particularly important in the beginning of an outbreak when it is unclear if mumps is the cause. If the patient has orchitis/oophoritis, mastitis, pancreatitis, hearing loss, meningitis, or encephalitis, collect a buccal swab specimen for rRT-PCR and a serum specimen for IgM detection, regardless of days since symptom onset.
Q: What does a mumps-negative rRT-PCR result mean?
A: Failure to detect mumps virus RNA by rRT-PCR in specimen from a person with clinically compatible mumps symptoms does not rule out mumps as a diagnosis. Successful detection of mumps virus depends primarily on the timing of collection and quality of the clinical sample. Vaccinated individuals may shed virus for a shorter period and might shed smaller amounts of virus, thus degradation of the sample has greater consequences for successful detection of virus. In outbreaks among two-dose vaccine recipients, mumps virus RNA was detected in specimen from 30%–71% of case-patients if the specimen were collected within 3 days following onset of parotitis. IgM was detected in 13 to 50% of these cases (Bitsko et al, 2008, Rota et al. 2009, Rota et al, 2013).
Q: What does a mumps-positive rRT-PCR result mean?
A: A positive rRT-PCR signal indicates the presence of mumps virus RNA in the patient sample. A positive rRT-PCR result provides laboratory confirmation of mumps infection in persons with symptoms consistent with mumps.
Q: Why doesn’t a negative rRT-PCR result and/or a negative IgM result rule out mumps as a diagnosis?
A: Successful detection of mumps virus or anti-mumps IgM antibody is dependent upon the timing of specimen collection and quality of the clinical sample. Vaccinated individuals may shed virus for a shorter period and may shed smaller amounts of virus. Viral RNA may not be detectable in specimen that have been collected, stored or shipped improperly. IgM may be transient or absent and therefore not detected.
Q: What is a mumps genotype?
Q: What mumps virus genotype has been commonly detected in the United States?
A: Since CDC initiated routine genotype surveillance for mumps in 2006, CDC and reference centers have detected mostly genotype G among people with mumps in the United States. A few of the other 11 genotypes were also detected, but they are usually associated with mumps importations into the United States and have not been associated with large outbreaks.
Mumps outbreaks are typically associated with only one genotype. There are no differences in the genotypes detected in vaccinated and unvaccinated people who have gotten mumps in the United States.
For more information, please see the “Genetic characterization of mumps viruses associated with the resurgence of mumps in the United States: 2015-2017” study.
Q: Are there small genetic changes that may be affecting our molecular diagnostics?
A: Viruses like mumps have very high rates of genetic change compared to other organisms. The CDC diagnostic assay, rRT-PCR, is designed to detect a specific sequence that is conserved in many mumps genotypes. If genetic changes occur in the sequence that is being detected by rRT-PCR, the assay may lose the ability to detect mumps virus with high sensitivity. CDC monitors the performance of the RT-PCR assay in collaboration with the four Vaccine Preventable Disease Reference Centers that are supported by the Association of Public Health Laboratories (APHL) and CDC. In addition, state laboratories performing the rRT-PCR assay to detect mumps may participate in a proficiency testing program that was developed by CDC and the Reference Centers and is managed by the Wisconsin State Laboratory of Hygiene.
Q: Would genetic changes that are occurring cause the vaccine not to work against certain strains of the virus?
A: Not all genetic changes result in a change in viral proteins. “Silent mutations” change the genetic sequence without changing the predicted protein. Viral proteins are recognized by the host’s immune system. There are several viral proteins, each containing many recognition sites. Therefore, it would take many genetic changes to change the viral proteins to the extent that they are no longer recognized by the immune response in vaccinated individuals. CDC is developing tests to monitor the ability of serum from vaccinated individuals to neutralize currently circulating strains.
Q: If a clinical sample is determined to be positive by rRT-PCR or by viral isolation at the state laboratory, is it necessary to send a sample to CDC or a VPD-RC for sequence analysis?
A: Laboratories are encouraged to send patient specimen from positive sporadic cases of mumps as well as representative specimen from an outbreak. The sequence of the mumps short hydrophobic (SH) gene is used to assign mumps viruses to one of 12 recognized genotypes. The sequence information may help to identify the source of the virus and can provide confirmation of suspected epidemiologic links. Specimen for genotyping can be sent to the CDC or the VPD-RCs.
During ongoing outbreaks, it is not necessary to obtain a genotype on every rRT-PCR positive specimen although it is recommended to obtain representative sequences on a weekly basis. However, if the outbreak spreads to another community, an effort should be made to obtain a genotype for specimens from the new outbreak. Additionally, specimens should be submitted for genotyping from confirmed cases that do not meet the case definition, that have an unusually long incubation period, or have travel history (domestic or international).
Q: Where can I find RT-PCR and real-time RT-PCR protocols for mumps?
A: The following updated protocols are available online:
- Real-time RT-PCR protocol, targeting the nucleoprotein (N) gene [3 pages] (Updated Aug 2021)
- Standard RT-PCR for mumps virus [6 pages] (Updated Jun 2021)
For more information on changes made to the protocols, please see “Would genetic changes that are occurring cause the vaccine not to work against certain strains of the virus?”
Q: Where do I obtain material for rRT-PCR and virus isolation controls?
A: CDC can provide a sample of synthetic RNA for real-time RT-PCR reactions (N gene) and for genotyping (endpoint) RT-PCR (SH gene). This material is ready to use in RT-PCR reactions. If laboratories would like to produce their own RNA specimen or require a positive control for virus isolation, viruses can be obtained from ATCC or BEI. Public health laboratories or laboratories affiliated with state public health laboratories may send requests for mumps RNA to Dr. Bettina Bankamp. A proficiency test for mumps RT-PCR can be obtained from the Wisconsin State Laboratory of Hygiene.
References
- Bitsko RH, Cortese MM, Dayan GH, Rota PA, Lowe L, Iversen SC, Bellini WJ. Detection of RNA of mumps virus during an outbreak in a population with a high level of measles, mumps, and rubella vaccine coverage. J Clin Microbiol 2008;46:1101–3.
- Briss PA, Fehrs LJ, Parker RA, Wright PF, Sannella EC, Hutcheson RH, Schaffner W. Sustained transmission of mumps in a highly vaccinated population: assessment of primary vaccine failure and waning vaccine-induced immunity. J Infect Dis 1994;169:77–82.
- Davidkin I, Jokinen S, Paananen A, Leinikki P, Peltola H. Etiology of mumps-like illnesses in children and adolescents vaccinated for measles, mumps, and rubella. J Infect Dis 3005;191:719–23.
- Gouma S, Vermeire T, Van Gucht S, et al. Differences in antigenic sites and other functional regions between genotype A and G mumps virus surface proteins. Sci Rep. 2018;8(1):13337.
- Gut JP, Lablache C, Behr S, Kirn A. Symptomatic mumps virus reinfections. J Med Virol 1995;45:17–23.
- Hatchette TF, Mahony JB, Chong S, LeBlanc JJ. Difficulty with mumps diagnosis: what is the contribution of mumps mimickers? J Clin Virol 2009; 46:381-3.
- Jin, L., et al., Genomic diversity of mumps virus and global distribution of the 12 genotypes. Rev Med Virol, 2015. 25(2): p. 85-101.
- Jin L, Feng Y, Parry R, Cui A, Lu Y. Real-time PCR and its application to mumps rapid diagnosis. J Med Virol 2007; 79:1761-7.
- Krause CH, Eastick K, Ogilvie MM. Real-time PCR for mumps diagnosis on clinical specimens–comparison with results of conventional methods of virus detection and nested PCR. J Clin Virol 2006; 37:184-9.
- Krause CH., Molyneaux PJ, Ho-Yen DO, McIntyre P, Carman WF, Templeton KE. Comparison of mumps-IgM ELISAs in acute infection. J Clin Virol 2007;38:153–6.
- L’Huillier AG, Eshaghi A, Racey CS, et al. Laboratory testing and phylogenetic analysis during a mumps outbreak in Ontario, Canada. Virol J 2018; 15:98.
- Maillet M, Bouvat E, Robert N, et al. Mumps outbreak and laboratory diagnosis. J Clin Virol 2015; 62:14-9.
- Narita M, Matsuzono Y, Takekoshi Y, Yamada S, Itakura O, Kubota M, Kikuta H, Togashi T. Analysis of mumps vaccine failure by means of avidity testing for mumps virus-specific immunoglobulin G. Clin Diagn Lab Immunol 1998;5:799–803.
- Nunn A, Masud S, Krajden M, Naus M, Jassem AN. Diagnostic Yield of Laboratory Methods and Value of Viral Genotyping during an Outbreak of Mumps in a Partially Vaccinated Population in British Columbia, Canada. J Clin Microbiol. 2018;56(5).
- Rota JS, Rose JB, Doll MK, McNall RJ, McGrew M, Williams N, Lopareva EN, Barskey AE, Punsalang Jr A, Rota PA, Oleszko WR, Hickman CJ, Zimmerman DM, Bellini WJ. Comparison of the sensitivity of laboratory diagnostic methods from a well-characterized outbreak of mumps in New York City in 2009. Clin Vaccine Immunol. 2013;20:391-6.
- Rota JS, Rosen JB, Doll MK, et al. Comparison of the sensitivity of laboratory diagnostic methods from a well-characterized outbreak of mumps in New York city in 2009. Clin Vaccine Immunol. 2013;20(3):391-396.
- Rota JS, Turner JC, Yost-Daljev MK, Freeman M, Toney DM, Meisel E, Williams N, Sowers SB, Lowe L, Rota PA, Nicolai LA, Peake L, Bellini WJ. Investigation of a mumps outbreak among university students with two measles-mumps-rubella (MMR) vaccinations, Virginia, September–December 2006. J Med Virol 2009;81:1819–25.
- Rubin SA, Qi L, Audet SA, et al. Antibody induced by immunization with the Jeryl Lynn mumps vaccine strain effectively neutralizes a heterologous wild-type mumps virus associated with a large outbreak. J Infect Dis. 2008;198(4):508-515.
- Sakata H, Tsurudome M, Hishiyama M, Ito Y, Sugiura A. Enzyme-linked immunosorbent assay for mumps IgM antibody: comparison of IgM capture and indirect IgM assay. J Virol Methods 1985;12:303–11.
- Sartorius B, Penttinen P, Nilsson J, Johansen K, Jönsson K, Arneborn M, Löfdahl M, Giesecke J. An outbreak of mumps in Sweden, February–April 2004. Euro Surveill 2005;10 (9):pii=559. Available at https://www.eurosurveillance.org/content/10.2807/esm.10.09.00559-enexternal icon
- Tan KE, Anderson M, Krajden M, Petric M, Mak A, Naus M. Mumps virus detection during an outbreak in a highly unvaccinated population in British Columbia. Can J Public Health 2011; 102:47-50.
- Vermeire T, Barbezange C, Francart A, et al. Sera from different age cohorts in Belgium show limited cross-neutralization between the mumps vaccine and outbreak strains. Clin Microbiol Infect. 2018.