Emergence of Localized Serogroup W Meningococcal Disease in the United States — Georgia, 2006–2016
Weekly / August 17, 2018 / 67(32);894–897
Ashley E. Moore, MS, MPH1; Jessica R. MacNeil, MPH2; Xin Wang, PhD2; Sandeep J. Joseph, PhD2; Lauren Lorentzson, MPH1; Stepy Thomas, MSPH3; Amy Tunali, MPH3; Tonia Parrott, PhD4; Monica M. Farley, MD5; Melissa Tobin-D’Angelo, MD1 (View author affiliations)
View suggested citationSummary
What is already known about this topic?
The incidence of meningococcal disease has been declining in the United States for decades, but Neisseria meningitidis serogroup W incidence has been increasing in countries around the world.
What is added by this report?
The incidence of Neisseria meningitidis serogroup W is increasing in Georgia. Although not associated with an outbreak, molecular testing indicated that the Georgia serogroup W isolates are all from the same clonal complex, CC11. This strain is associated with an increased morbidity and mortality which could have severe implications.
What are the implications for public health practice?
The collection and testing of meningococcal isolates for serogroup and strain information is important to monitor changes and emergence of previously underrepresented serogroups.
Several countries in Europe and Australia are reporting an increasing incidence of Neisseria meningitidis serogroup W (NmW) as a consequence of the rapid expansion of a single NmW clone belonging to clonal complex 11 (1–5). Because this clone is reported to be associated with more severe disease, unusual clinical presentations, and a high case fatality ratio (CFR), it is considered a hypervirulent strain (1,6). In the United States, NmW accounts for approximately 5% of meningococcal disease reported each year, and this proportion has remained stable for several years (7). However, localized increases in NmW have been reported, most notably in Florida during 2008–2009 (8). In Georgia, NmW accounted for only 3% of meningococcal disease cases reported during 2006–2013; however, between January 2014 and December 2016, 42% of all reported cases were NmW. Surveillance data from Georgia were analyzed to describe the epidemiology and clinical characteristics of NmW cases, and whole-genome sequencing of NmW isolates was performed for comparison with NmW strains circulating in the United States and worldwide. These data indicate that the U.S. NmW strains might have evolved from the same ancestor as the hypervirulent strain that is circulating globally. Genetic analysis demonstrates that these strains are closely related, which would suggest that genetic variation led to the rise of different strains from the same ancestor. Given the recent global expansion of this potentially hypervirulent NmW lineage, clinicians and public health officials need to remain vigilant in obtaining isolates to monitor changes in circulating strains.
A case of meningococcal disease was defined as laboratory-confirmed N. meningitidis isolated from a normally sterile body site, reported to the Georgia Department of Public Health (DPH) during 2006–2016. A comprehensive case report form, developed for the Emerging Infections Program’s Active Bacterial Core surveillance (9), was used to abstract case medical record data, including demographic and clinical information. Clinical syndromes (e.g., bacteremia, meningitis, pneumonia) were not mutually exclusive; a patient could have multiple syndromes simultaneously. For statistical comparisons, Fisher’s Exact and Student’s t-Test statistics were calculated; p-values <0.05 were considered statistically significant.
All N. meningitidis isolates were requested for serogroup typing at the Georgia Public Health Laboratory as part of Active Bacterial Core surveillance. The isolates were then forwarded to CDC for serogroup confirmation and further molecular characterization using whole genome sequence analysis. The phylogenetic analysis included 18 NmW isolates collected in Georgia during 2012–2016, isolates from other states collected through routine surveillance, and the genome sequences of the global strains, obtained from the Bacterial Isolate Genome Sequence Database of PubMLST,* public databases for molecular typing and microbial genome diversity.
During 2006–2016, a total of 178 meningococcal disease cases were reported to DPH, including 158 (89%) with isolates available for serogroup typing. The 20 patients without an isolate available for serogroup typing were excluded from the analysis; these patients did not differ significantly by race, age, or sex from those with a known serogroup.
Overall, 21 (13%) NmW cases and 137 (87%) N. meningitidis non-serogroup W (non-NmW) cases were identified; the proportion of NmW cases increased from 0% in 2013 to 47% in 2016 (Figure 1). No epidemiologic links were identified among the patients with NmW disease, although 70% of NmW cases reported since 2006 were concentrated geographically in northern Georgia.
Among 21 patients with NmW disease, 14 (68%) were male compared with 74 (54%) patients with non-NmW disease; however, this difference was not statistically significant (Table). The median age of patients with NmW disease (34 years) was significantly higher than that of patients with non-NmW disease (26 years) (p = 0.005); 90% of patients with NmW were aged ≥18 years compared with 61% of patients with non-NmW disease. Data on admission to an intensive care unit (ICU) has been collected for all meningococcal disease cases since 2010; from 2010–2016, a similar percentage of patients with NmW disease and non-NmW disease were admitted to an ICU (56% and 54%, respectively). The CFR was higher for patients with NmW (24%) than for patients with non-NmW disease (15%); however, the numbers are small and were not statistically significantly different.
Bacteremia was reported in 50% of NmW and 35% of non-NmW cases, and meningitis accounted for less than 40% of infections in both groups. Although not collected systematically for all meningococcal disease cases in Georgia, it was noted in medical records that nine (41%) NmW patients during 2014–2016 reported gastrointestinal (GI) symptoms, such as diarrhea and vomiting, to their providers.
Eighteen (86%) NmW isolates belonged to clonal complex 11 (CC11); 17 of these were sequence type 11 (ST-11), and one, ST-10826, was a new sequence type. Pairwise comparison, a process of comparing any two sequences for genetic differences, indicated the difference between each pair of the 18 Georgia isolates ranged from 0–63 single nucleotide polymorphisms. The 17 ST-11 isolates from Georgia were more similar to each other than to isolates tested from other states (California, Florida, Ohio, and Texas) (Figure 2). Overall, the U.S. NmW CC11 isolates were more similar to strains from South America and Europe (six from the United Kingdom) than to those from Africa (Figure 2).
Discussion
Georgia experienced an increase in NmW disease during 2014–2016, compared with 2006–2013, which was associated with the emergence of a CC11 NmW strain that is different from the CC11 NmW strains from other U.S. states. Phylogenetic comparison of the Georgia and other U.S. CC11 NmW strains with global isolates indicates that these U.S. strains might have evolved from a clone previously observed in South America, which is also an ancestor of the hypervirulent United Kingdom strain that has emerged in Europe and Australia (5).
In contrast to other published reports, this analysis did not identify significant differences in CFR or clinical presentation of patients with NmW disease compared with those with non-NmW disease. However, there was a slightly higher frequency of ICU admission and higher CFR in patients with NmW disease, which are consistent with a report from the United Kingdom that found that older children and adults with NmW disease were more likely to be admitted to the ICU (1). In addition, many NmW patients in the United Kingdom had predominantly GI symptoms, diarrhea in particular, which reportedly led to initial misdiagnoses and delays in provision of appropriate care (6). Although 41% of the Georgia NmW patients did report GI symptoms, information on these symptoms was not systematically collected on all meningococcal cases for comparison.
In the United Kingdom, the emergence of cases caused by the hypervirulent ST-11 strain initially began in adults but quickly extended to other age groups; during 2013–2014, this ST-11 strain accounted for nearly all NmW cases in persons aged 5–64 years and a high proportion of NmW cases in other age groups (1). This is of interest because in this analysis 90% of NmW cases occurred in persons aged ≥18 years; therefore, surveillance data will need to be monitored closely for future shifts in the age distribution of NmW cases.
The findings in this report are subject to at least three limitations. First, cases of N. meningitidis are rare, and thus performing sufficiently powered statistical tests of significance on the data are difficult. Second, serogroup W cases only make up 5% of reported meningococcal cases each year in the United States, and as a result, the comparison group for isolates within the United States is limited. Finally, clinical presentation and symptoms were not collected systematically for all N. meningitidis cases, which precluded direct analysis of Georgia data and comparison with data from other countries.
Although the numbers in this study are small, this report provides description of the NmW clone that has emerged in Georgia and its associated cases. The DPH will continue to monitor and follow up on all patients with meningococcal disease to collect clinical information and isolates to determine whether the trend of an increasing proportion of NmW cases continues. Clinicians and public health officials need to remain vigilant in obtaining isolates from all cases of meningococcal disease to monitor changes in circulating strains over time, and also remain aware of the potential for atypical clinical presentations that might not be indicative of meningococcal disease to prevent delays in treatment that could result in unnecessary morbidity and mortality.
Acknowledgments
Active Bacterial Core Surveillance program, CDC; Emerging Infections Program, CDC; hospitals, laboratories, infection preventionists, and local health departments within the state of Georgia.
Conflict of Interest
No conflicts of interest were reported.
Corresponding author: Ashley E. Moore, Ashley.Moore@dph.ga.gov, 404-657-2643.
1Georgia Department of Public Health; 2National Center for Immunizations and Respiratory Diseases, CDC; 3Atlanta Research Education Foundation, Veterans Association Medical Center, Atlanta, Georgia; 4Georgia Department of Public Health Laboratory, Decatur, Georgia; 5Division of Infectious Diseases, Emory University School of Medicine, Atlanta, Georgia.
References
- Ladhani SN, Beebeejaun K, Lucidarme J, et al. Increase in endemic Neisseria meningitidis capsular group W sequence type 11 complex associated with severe invasive disease in England and Wales. Clin Infect Dis 2015;60:578–85. CrossRef PubMed
- Hong E, Barret AS, Terrade A, et al. Clonal replacement and expansion among invasive meningococcal isolates of serogroup W in France. J Infect 2018;76:149–58. CrossRef PubMed
- Martin NV, Ong KS, Howden BP, et al. ; Communicable Diseases Network Australia MenW Working Group. Rise in invasive serogroup W meningococcal disease in Australia 2013-2015. Commun Dis Intell Q Rep 2016;40:E454–9. PubMed
- Carville KS, Stevens K, Sohail A, et al. Increase in meningococcal serogroup W disease, Victoria, Australia, 2013–2015. Emerg Infect Dis 2016;22:1785–7. CrossRef PubMed
- Lucidarme J, Scott KJ, Ure R, et al. An international invasive meningococcal disease outbreak due to a novel and rapidly expanding serogroup W strain, Scotland and Sweden, July to August 2015. Euro Surveill 2016;21:30395. CrossRef PubMed
- Campbell H, Parikh SR, Borrow R, Kaczmarski E, Ramsay ME, Ladhani SN. Presentation with gastrointestinal symptoms and high case fatality associated with group W meningococcal disease (MenW) in teenagers, England, July 2015 to January 2016. Euro Surveill 2016;21:30175. CrossRef PubMed
- MacNeil JR, Blain AE, Wang X, Cohn AC. Current epidemiology and trends in meningococcal disease—United States, 1996–2015. Clin Infect Dis 2018;66:1276–81. CrossRef PubMed
- Doyle TJ, Mejia-Echeverry A, Fiorella P, et al. Cluster of serogroup W135 meningococci, southeastern Florida, 2008-2009. Emerg Infect Dis 2010;16:113–5. CrossRef PubMed
- Schuchat A, Hilger T, Zell E, et al. ; Active Bacterial Core Surveillance Team of the Emerging Infections Program Network. Active bacterial core surveillance of the emerging infections program network. Emerg Infect Dis 2001;7:92–9. CrossRef PubMed
 FIGURE 1. Meningococcal disease cases, by serogroup type — Georgia, 2006–2016
FIGURE 1. Meningococcal disease cases, by serogroup type — Georgia, 2006–2016
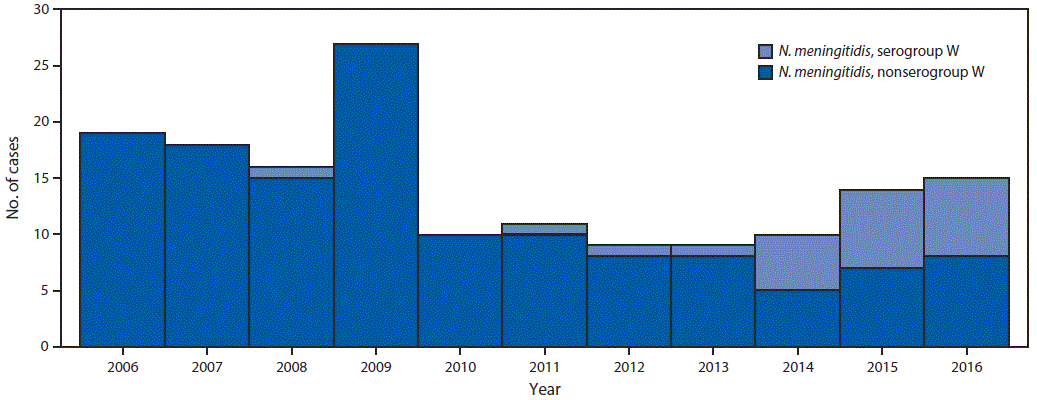
Abbreviation: N. meningitidis = Neisseria meningitidis.
The figure above is a histogram showing meningococcal disease cases, by serogroup type, in Georgia during 2006–2016.
Abbreviations: ICU = intensive care unit; NmW = N. meningitidis serogroup W; Non-NmW = N. meningitidis nonserogroup W.
* Unknowns excluded from the table and the denominator calculations.
† Other infections not listed include pneumonia, septic arthritis, puerperal sepsis, cellulitis, epiglottitis, and supraglottitis. With the exception of bacteremia only, a patient might have multiple types of infections simultaneously.
§ This variable was not collected before 2010.
 FIGURE 2. Whole genome maximum likelihood phylogeny* of N. meningitidis serogroup W clonal complex 11 isolates from Georgia, other U.S. states, Europe, Africa, and South America, 2006–2016
FIGURE 2. Whole genome maximum likelihood phylogeny* of N. meningitidis serogroup W clonal complex 11 isolates from Georgia, other U.S. states, Europe, Africa, and South America, 2006–2016
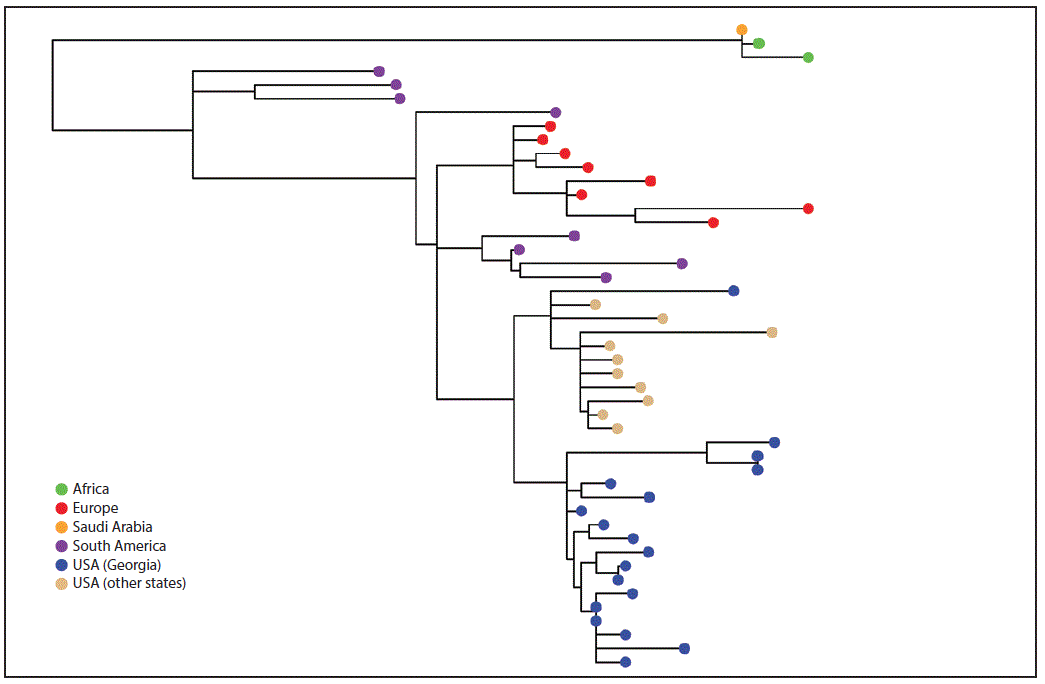
* Branch length is related to the number of nucleotide substitutions. The more substitutions an isolate has, the longer its branch will be. More evolved strains will be further from their ancestor.
The figure above is phylogenetic tree showing the whole genome maximum likelihood phylogeny of N. meningitidis serogroup W clonal complex 11 isolates from Georgia, other U.S. states, Europe, Africa, and South America during 2006–2016.
Suggested citation for this article: Moore AE, MacNeil JR, Wang X, et al. Emergence of Localized Serogroup W Meningococcal Disease in the United States — Georgia, 2006–2016. MMWR Morb Mortal Wkly Rep 2018;67:894–897. DOI: http://dx.doi.org/10.15585/mmwr.mm6732a5.
MMWR and Morbidity and Mortality Weekly Report are service marks of the U.S. Department of Health and Human Services.
Use of trade names and commercial sources is for identification only and does not imply endorsement by the U.S. Department of
Health and Human Services.
References to non-CDC sites on the Internet are
provided as a service to MMWR readers and do not constitute or imply
endorsement of these organizations or their programs by CDC or the U.S.
Department of Health and Human Services. CDC is not responsible for the content
of pages found at these sites. URL addresses listed in MMWR were current as of
the date of publication.
All HTML versions of MMWR articles are generated from final proofs through an automated process. This conversion might result in character translation or format errors in the HTML version. Users are referred to the electronic PDF version (https://www.cdc.gov/mmwr) and/or the original MMWR paper copy for printable versions of official text, figures, and tables.
Questions or messages regarding errors in formatting should be addressed to mmwrq@cdc.gov.