Considerations and Resources for Conducting an On-Farm Environmental Investigation
Intended Audience: This information can be used by the produce industry to conduct on-farm environmental investigations of foodborne pathogen contamination in the growing environment. This contamination could be detected during routine environmental monitoring or following a suspected health risk.
A retrospective investigative method used to determine how the root cause of a trigger event occurred and provide information for use in determining what actions can be taken to eliminate the root cause and to prevent recurrence of the trigger event.
An investigation of the environment and associated factors contributing to the presence, growth, or transport of elevated or harmful microbes that represent a potential public health risk. This includes evaluating the potential contamination sources, the environmental controls in place, and the ways that people and animals interact.
The objectives of this document are to provide considerations for the benefits (both short- and long-term), challenges, and limitations of doing an on-farm environmental investigation and resources for conducting an environmental investigation as a component of a root cause analysis (RCA) in a produce growing environment. The examples used as the antecedent events are Shiga toxin-producing Escherichia coli (STEC) contamination in the growing environment or elevated Escherichia coli (E. coli) levels in agricultural water used for fresh produce production. While this document provides considerations and resources for conducting an environmental investigation, it does not provide a step-by-step protocol, as all scenarios will be unique. A produce growing environment presents unique challenges for conducting an environmental investigation that can limit the success of finding the root cause or mitigating its impact. However, collecting and analyzing environmental data and data about on- and off-farm practices can provide invaluable insights into potential foodborne pathogen (i.e., disease-causing microorganism) contamination risks and prevention strategies.
Table of Contents
- Overview and role of on-farm environmental investigations
- Challenges, limitations, and benefits of environmental investigations
- Resources for environmental investigations
Overview and Role of an On-Farm Environmental Investigation
An RCA may be conducted in response to a foodborne outbreak caused by a contaminated product, a product recall from pathogen detection during finished product testing, detection of a pathogen on the pre-harvest product or within the growing environment, or elevated E. coli levels in agricultural water. In these instances, the RCA may warrant an environmental investigation in addition to the other investigative steps conducted during the RCA.
Factors to consider:
- Water quality and use
- Growing practices
- Adjacent land use
- Wildlife proximity
- Weather patterns
- Regulatory changes
- Process changes
The objective of an environmental investigation is to collect the data needed to identify the root cause (i.e., how and why the problematic event happened). In the example of on-farm STEC contamination, the root cause would be the fecal source and mode(s) of contamination. However, a growing environment is a dynamic and complex system that is affected by both the direct actions of the grower and the indirect (and potentially unknown) influences of the surrounding environment. Therefore, establishing a hypothesis about which factors may have contributed to the problem is a critical component guiding an environmental investigation. Hypothesis generation can be challenging without enough information to identify suspected contributing factors. The practice of ongoing, systematic environmental monitoring and analysis is recommended because it can aid in identifying trends, patterns, or associations that can point to suspected factors which may contribute to on-farm contamination.
Challenges, Limitations, and Benefits of an Environmental Investigation
The goals of any RCA effort are to identify the root cause(s) (the why and how) of contamination in order to identify actions needed to eliminate the problem and prevent it from happening again. However, conducting an environmental investigation as part of an RCA presents unique challenges and limitations.
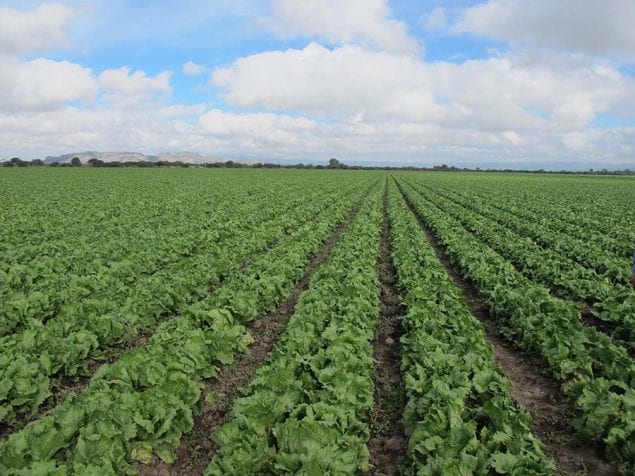

Identification of the root cause may not be possible because:
- Unknown number of potential contributing factors to investigate. In an open environment, the number of contributing factors could be numerous, so identifying a root cause is nearly impossible if every contributing factor must be investigated. An environmental investigation is much more likely to be successful if you have already identified suspected contributing factors with which to form a hypothesis for investigation – such as from an outbreak investigation traceback of common case food exposures, a product recall event, or an ongoing analysis of routine environmental monitoring efforts.
- The investigation scope often extends off-farm. Because of the influences of the surrounding environment, it is probable that an environmental investigation will extend beyond the boundaries of the produce growing operation. This could require cooperation of external firms and municipalities and data sharing between these entities.
- Conditions may have changed since the contamination event. The root cause of on-farm contamination may be transient and difficult to identify retrospectively, and the more time that has elapsed between the event and initiation of an RCA, the lower the chance of identifying the root cause. In this kind of complex setting, the effort required to get to the root cause may be time- and resource-prohibitive.
Preventing contamination and eliminating health risk may not be possible because:
- The root cause can consist of naturally occurring elements and modes of environmental contamination that are often in flux. A growing environment is an open system affected by influences of the surrounding environment, where zoonotic pathogens such as STEC, which may be shed in animal feces, are naturally present. Pathogen occurrence and concentration in certain settings may fluctuate due to naturally occurring and complex environmental factors. The pathogen may also be naturally persistent in the environment.
- Corrective actions may not be able to control or mitigate the problem. Environmental factors contributing to an event may be identifiable, but it may not be possible to control those factors to mitigate or remediate the problem to prevent future events (e.g., close proximity of animal operations, reliance on surface water for produce irrigation).
Despite these challenges and limitations, conducting an environmental investigation may still prove to be beneficial to improving food safety since it may identify issues not initially considered. Regardless of the outcome of an environmental investigation, an RCA provides an opportunity to learn more about operating procedures and processes and may reveal unanticipated issues that need to be addressed or identify areas where improved communication or additional training, documentation, data collection, or testing is warranted. While corrective or preventive actions may not be able to control or mitigate every on-farm risk or contributing factor identified, the information acquired from an environmental investigation may inform whether controllable practices should be modified or monitored differently. In addition to the ongoing data analysis from monitoring efforts, the data collected during environmental investigations can aid in uncovering associations, patterns, and trends associated with the risk of potential on-farm pathogen contamination. This information can be used to generate hypotheses to guide future RCAs or larger-scale evaluations or to spur the implementation of new evidence-based safety standards and prevention efforts.
Resources for Environmental Investigation
Below are resources for conducting an environmental investigation using on-farm STEC contamination or elevated E. coli levels in agricultural water as example scenarios. A brief overview of environmental sampling and testing concepts is provided along with examples of methods to consider.
Environmental Investigation Team
In addition to the industry and food safety members of the overall investigation team, the environmental investigation team should include an environmental microbiologist and/or environmental engineer. These subject matter experts (SMEs) can aid in hypothesis generation and are crucial for designing the site- and contamination-specific sampling plans and choosing the appropriate sample collection and testing methods to address the hypothesis being investigated. University cooperative extension offices or state agricultural assistance programs may have the applicable SMEs or be able to help identify those who can assist in your investigation.1,2 Additional resources may also be available from industry trade organizations.
Sampling and Analysis Plan
The sampling and analysis plan will be dependent on the problem, setting, and root cause hypotheses being tested and should identify:
- The types of samples to collect. The types of samples collected (e.g., water, soil, sediment, swabs, product) may provide information about potential pathogen contamination routes, reservoirs or environmental niches, persistence, and extent of contamination.
- Where to collect samples. The chosen sample sites (e.g., irrigation water source, water conveyance systems and holding tanks, water used by harvesting equipment, surfaces of equipment/tools) should include locations to test the investigation hypothesis (suspected fecal source and modes of contamination) and should include sites expected to be positive and negative in order to test the hypothesis.
- How many samples to collect. The samples collected should ensure representative sampling of potential pathogen sources and modes of contamination. If an investigation warrants repeated sampling from specific sites to investigate time-dependent variables associated with contamination, the frequency of sampling should also be determined.
- What to test the samples for. The samples should be tested for physicochemical parameters and microbial targets that test the investigation hypothesis or provide information about changes or trends in the sampling location (if historical information is available).
Sample Collection Methods
There are limitations to the conclusions that can be drawn from microbial testing of environmental samples. Detection verifies the presence of a pathogen or analyte at the time of sample collection, as well as the concentration if a quantitative test was conducted. Conversely, not detecting the pathogen or analyte indicates that the specific sample volume did not contain the target at the time of collection or that the concentration was below the detection limit of the test, but does not necessarily rule out the presence of the pathogen or analyte in the larger environment from which the sample was taken. Testing cannot rule out the presence of the target in the sampling environment, nor can it rule out the presence of the target prior to sample collection or at any point in the future.
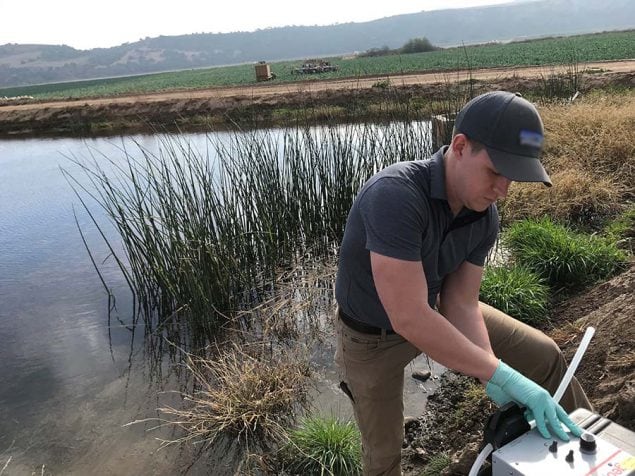

Water
Water can be an effective vehicle of large-scale on-farm contamination. Sample collection methods typically used for routine testing of water for generic E. coli may not be appropriate for use during an RCA environmental investigation. Small-volume samples (100 mL – 1 L), such as those collected for routine water monitoring, are appropriate for fecal indicator (e.g., E. coli) testing or for measures such as water quality parameters.1 However, many pathogens typically shed in feces are likely present in water samples at concentrations too low to be detected in ≤1-liter volumes due to various environmental factors such as dilution and decay. Because of this, collection of large-volume samples is often required for pathogen detection in the environment. Collection of large-volume samples reduces the microbe detection limit to well below 1 microbe per 100 mL or 1 L.
A commonly used method for collecting large-volume water samples is dead-end ultrafiltration (DEUF), which is a robust method for co-collection of microbes including bacteria, parasites, and viruses.2,3 During the DEUF procedure, water is pumped through a kidney dialysis filter at the sampling site, which traps microbes and filters out the water. Ultrafilters are then shipped to a laboratory and backflushed to recover the concentrated microbes. DEUF has recently been adopted by the U.S. Food and Drug Administration (FDA) for Cyclospora detection from agricultural water.4 The U.S. Environmental Protection Agency (USEPA) and CDC joint protocol provides instruction for collection of water samples by DEUF for detection of various pathogens, including STEC.2 For surface water samples, filtration of a minimum of 10 liters up to 50 liters is recommended depending on water quality and clarity. For ground water or well water samples, filtration of at least 100 liters is recommended. The SME should determine the desired sample volume based on water type and quality, desired detection limit, and time constraints.
Moore swabs may be used if a composite water sample over time is desired. Moore swabs consist of a compressed or folded cheesecloth in a permeable container or cartridge, which is placed in a body of water for hours to days to allow water to passively flow through the cheesecloth.5
Submerged sediment and biofilm
Many zoonotic pathogens can survive and persist in submerged sediment or in biofilms for prolonged periods. Collection of submerged sediment or biofilm swabs may be paired with collection of bulk water to compare the potential of historic contamination with current contamination. There are numerous methods for collection of sediment,6 but for microbial testing of submerged sediment, collection of the first few inches of sediment with a sterile scoop into a sterile collection vessel is sufficient. Swab collection methods vary based on the target microbe, type of surface, and whether quantification is required.7 For microbial testing during an environmental investigation, cellulose sponge swabs are effective for many types of environmental surfaces. Composite sample collection (combining multiple samples into one container to test as one sample) may be a useful way of obtaining a representative sample of a large area under investigation.
Pre-harvest product
Whether pre-harvest product testing is warranted for an environmental investigation depends on the hypothesis being tested and whether product testing will aid in identifying the root cause. Methods for collection of pre-harvest product for routine monitoring are also appropriate for collection during an environmental investigation. Composite sample collection may be a useful way of collecting a representative sample of the area under investigation.
Animal feces
Cattle and wildlife are natural reservoirs of zoonotic foodborne pathogens such as STEC and Salmonella. Thus, for these zoonotic pathogens, animal feces are the source of the pathogen’s transmission to the environment. Local animal feces should also be collected to evaluate the performance of any microbial source tracking assays used during the investigation. Collecting fecal samples of suspected animal sources may require access to neighboring businesses and permission to collect samples. Animal care and human safety should be considered when collecting a fecal sample directly from an animal. Collecting a fecal sample from the environment should be done in a manner that minimizes collection of the surrounding environment.7 Composite collection of animal feces may provide more coverage of fecal material and increase likelihood of pathogen detection if shedding is inconsistent and/or intermittent within a herd or flock.
Surface swabs
Zoonotic pathogens may be able to survive on surfaces for prolonged periods under favorable environmental conditions, such as surfaces that are wetted frequently. The surfaces chosen for sampling should have the potential to aid in root cause determination, such as interior surfaces of water storage or transport vessels. Similar to the methodology used for submerged biofilm, cellulose sponge swabs can be used to collect surface biofilm from many types of surfaces.7 Composite collection of surface swabs may provide more coverage of surfaces and increase likelihood of pathogen detection.
Soil
Many zoonotic pathogens can survive and persist in soil for prolonged periods under favorable environmental conditions. Whether soil testing is warranted for an environmental investigation depends on the hypothesis of the investigation and whether soil testing will aid in identifying the root cause. If so, collection of soil samples should be focused on only the area(s) of interest. While there are many methods for collecting soil samples for microbial testing during an environmental investigation,7,9,10 collection of the first few inches of soil with a sterile scoop into a sterile collection vessel is likely sufficient for evaluating recent contamination events. Composite collection of soil may provide more coverage of the area and increase likelihood of pathogen detection. Drag swabs have been used for sampling of poultry litter for Salmonella detection and have been used as an alternative composite sampling method to grab sample collection of sub-surface soil or soil cores.11,12 However, data are not available on the performance of drag swabs for recovery of STEC from soil.
Bioaerosols
Bioaerosols are airborne collections of biological material including suspended soil in the form of dust and could be an effective mode of contamination to a large amount of produce. Bioaerosol sampling for microbial detection utilizes liquid impingers,7 in which air is pulled through the impinger and microbes are trapped in a liquid medium or broth. Because bioaerosol presence in the environment is likely to be intermittent, collection of bioaerosol samples over time and at different wind velocities and directions may be warranted.
Resources:
- Standard Methods for the Examination of Water and Wastewater
- USEPA/CDC Protocol for Collection of Water Samples for Detection of Pathogens and Biothreat Agents
- Water Sampling and Processing Techniques for Public Health-Related Microbes. In Manual of Environmental Microbiology. Washington, D.C.: ASM Press; 2020.
- BAM Method 19C. Dead-end Ultrafiltration for the Detection of Cyclospora cayetanensis from Agricultural Water [PDF – 24 pages]
- Modified Moore swab optimization and validation in capturing E. coli O157:H7 and Salmonella enterica in large volume field samples of irrigation water [PDF – 9 pages]
- Ohio EPA Sediment Sampling Guide and Methodologies [PDF – 35 pages] (2nd Edition)
- USEPA Sample Collection Information Document for Pathogens–Companion to Selected Analytical Methods for Environmental Remediation and Recovery (SAM) 2017 (soil/swab/bioaerosol sample collection)
- USDA Cattle Fecal Collection Instructions [PDF – 7 pages] (fecal sample collection)
- USEPA/USGS Sample Collection Protocol for Bacterial Pathogens in Surface Soil
- USEPA Soil Sampling
- Landscape and Meteorological Factors Affecting Prevalence of Three Food-Borne Pathogens in Fruit and Vegetable Farms (example of drag swab use)
- Environmental Assessment of Factors Potentially Contributing to the Contamination of Romaine Lettuce Implicated in a Multi-State Outbreak of E. coli O157:H7 (Memorandum to the File on the Environmental Assessment)
Sample Testing Methods
Environmental samples should be tested for physicochemical parameters and microbial targets that test the investigation hypothesis or provide information about changes or trends in the sampling location (if historical information is available). The sample types described above can be analyzed by either molecular [e.g., polymerase chain reaction (PCR)] or culture methods. The laboratory will use a test method that is appropriate for detection of the specific analyte or pathogen from the specific environmental sample type.1,2 Using methods designed for environmental samples ensures that meaningful sample volumes can be analyzed. Environmental testing methods are also optimized to detect low levels of the pathogen or analyte of interest from a complex sample type with high numbers of competing or closely related organisms. The environmental investigation team should consider whether a quantitative result is desired; however, for most environmental investigations, a presence/absence result is sufficient. The laboratory should have proficiency in the selected methods and documentation of precision and accuracy of the method performance (also referred to as method recovery efficiency), along with known method sensitivity, specificity, and limit of detection. In addition to the test results, the laboratory should provide the results of all positive and negative controls, method blanks, and inhibition controls.
The following are considerations for environmental investigation testing tools.
Pathogen assays
Pathogen testing in an environmental investigation that is part of a root cause investigation is intended to test the possible root cause(s) of on-farm contamination (pathogen source and modes of environmental contamination). Culture testing methods are specific to the target or pathogen of interest and the sample type. In the case of an environmental investigation due to STEC contamination, a method capable of detecting a broad range of STECs, such as FDA’s BAM Chapter 4A: Diarrheagenic Escherichia coli,3 may be warranted. If the specific STEC serogroup or serotype is known, such as E. coli O157:H7, a more targeted method, such as USEPA’s Standard Analytical Method for E. coli O157:H7,4 may be more successful for recovery from water, sediment, and other environmental matrices. While these methods are optimized for detection from food and water, respectively, other environmental sample types may be used as inputs, provided the laboratory evaluates method performance data for new sample types. The SME microbiologist should work with the laboratory to choose the best method for the pathogen target and the format of the sample type that is an appropriate input to the method.
Example Bacterial Typing Nomenclature – E. coli O157:H7
Pathotype STEC
Serogroup O157
Serotype O157:H7
Genotype WGS-type
Culture testing for an environmental investigation is recommended for a pathogen such as STEC. Culture methods include steps for characterizing the detected isolates to the serotype level using phenotypic or genetic methods. Following culture, PCR assays are run to confirm pathogen detection and identify the virulence genes in the isolates. However, this information alone does not provide the resolution needed to compare pathogen isolates detected from different samples during an environmental investigation. Because there may be multiple STEC types shed by a variety of animal hosts and present in the environment, it will likely be necessary to characterize the STEC isolates found in different samples to the serotype or genotype level to compare whether the isolates detected are of the same serotype and closely related genetically. If this high level of discrimination is not achieved, it may hinder the ability to identify the root cause or it may misidentify the root cause. Genomic sequencing methods such as whole genome sequencing (WGS) can be used to evaluate the genetic relatedness between isolates, which is the gold standard for providing the highest level of isolate discrimination.
Whole genome sequencing–based subtyping methods
The genome of an organism is made up of a sequence of DNA or RNA bases, and the number of differences in DNA or RNA bases between organisms is a measure of how closely related they are to one another. Whole genome sequencing is a laboratory procedure that determines the sequence of DNA bases in the genome of an organism.5 WGS-based subtyping analysis approaches include high-quality single nucleotide polymorphism typing (hqSNP), core genome multilocus sequence typing (cgMLST), and whole genome MLST (wgMLST) analyses. These methods determine the unique sequence of the isolate genome. WGS data can also be used to determine serogroup, serotype, and presence of specific genes of interest like virulence and antimicrobial resistance genes. Using DNA sequence analysis tools, the genome sequences from multiple isolates can be compared to determine how closely related they are, which reveals whether the pathogens detected from samples during an environmental investigation are sufficiently similar genetically or may have originated from the same fecal source. Genome sequences can be submitted to public databases for comparison to food, environmental, and human clinical isolates, such as the National Center for Biotechnology Innovation (NBCI) Pathogen Detection database.6 Bioinformatic expertise will be needed to analyze and interpret genomic sequence data.
Microbial source tracking
Testing for animal-specific fecal markers by microbial source tracking (MST) can be used to assess the contribution of fecal contamination to the environment. Fecal MST markers are detected using PCR to identify gene targets specific to the animal of interest, such as ruminants or birds.7,8 Detection of animal-specific fecal MST markers can indicate which animals may be contributing to contamination of the environment or, conversely, which animals may have picked up the contamination from the environment. Detection of animal-specific MST markers cannot indicate whether zoonotic pathogens are also present in the sample, but it does indicate a risk of pathogen contamination associated with feces from that animal and can direct an environmental investigation to the potential animal source or mode of transmission of the pathogen.
Environmental data
Environmental data from sampling sites or physicochemical data from the sample provide important corollary information to pathogen and MST data. Environmental parameters within the sampling site that should be considered for collection may include air temperature, precipitation, and wind speed and direction. Physicochemical parameters of a sample to collect may include temperature, pH, conductivity, salinity, and disinfectant residual if applicable. In addition, metadata about the collection event should be recorded, such as a unique sample identifier, GPS coordinates and sample location details, person(s) collecting the sample, sample type and method of collection, volume or amount collected, and date and time collected.
Resources:
- Food Safety Resource Clearinghouse National Water Quality Testing Labs Map
- Arizona Laboratories Conducting Soil, Plant, Feed or Water Testing [PDF – 2 pages]
- BAM Method 4A. Diarrheagenic Escherichia coli
- USEPA Standard Analytical Method for E. coli O157:H7
- What is whole genome sequencing?
- National Center for Biotechnology Innovation (NBCI) Pathogen Detection database
- USEPA Microbial Source Tracking Guide Document [PDF – 133 pages]
- Microbial source tracking markers for detection of fecal contamination in environmental waters: relationships between pathogens and human health outcomes
Data Management and Analysis
Data collected during an on-farm RCA environmental investigation should be entered into a data management tool such as a spreadsheet or database to maintain a historical record that can be referenced later. Data should be maintained in accordance with applicable quality management guidelines. Include data collected from all parts of the investigation, including the environmental investigation and any relevant environmental or operational data collected during the RCA about the time frame of interest. Each row of data should correspond to a single sample. Columns should correspond to the data elements that were collected during the investigation (e.g., type of sample, collection method, collection date, testing laboratory, test result).
Example spreadsheet used to maintain the data collected during sampling and testing
Analyses on the information collected during the RCA, alongside historical information, may be performed depending on the type of resulting data. This may include comparing collected information to set standards or information from previous investigations to identify changes. For most RCAs, statistical analyses may not be needed; however, when multiple samples on the same source are taken over time, statistical analyses can help to identify patterns and remove the “noise” of expected fluctuations. Review the data using visual inspection via charts, graphs, and summary tables to identify baseline trends and anomalies in the data.
Summarizing the environmental investigation methodology, results, and analyses can be useful to encapsulate the investigation for future reference and dissemination to stakeholders and the food safety community.
Resources:
- Widely available spreadsheet software, such as Excel or Google sheets, are sufficient for maintaining datasets that are not large in size and for visualization of the data
- Epi InfoTM – General use data collection and management software originally developed at CDC for outbreak investigations, but adaptable to other needs and free to the public. Includes a suite of tools including statistics, tables, graphs, and mapping of locations.
- Checklist for Creating a Data Management Plan
- SheetHacks: Discover the best tips and tricks for Google Sheets and Excel
- Tutorials developed by Duke University to use Excel for data analysis and visualization: