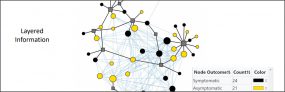
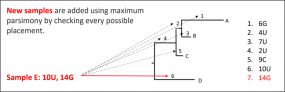
Module 3.4 – Walking through Nextstrain trees
Overview: This module demonstrates how to navigate through Nextstrain phylogenetic trees using various functionalities such as filtering, zooming, coloring and labeling to further analyze SARS-CoV-2 genomic epidemiological data. Nextstrain provides a list of SARS-CoV-2 resourcesexternal icon and publicly available analyses to help explore their tool.
Posted: 03/12/21
Presenter: Krisandra Allen, MPH, MB(ASCP)CM (view bio)
Molecular Epidemiologist
Washington State Department of Health
Further Reading:
- Nextstrain SARS-CoV-2 resources.external icon
Nextstrain, updated 2021.
Additional Resources:
- Auspice: An open-source interactive tool for visualising phylogenomic data.external icon
Nextstrain, updated 2020. - SARS-CoV-2 Sequencing for Public Health Emergency Response, Epidemiology and Surveillance (SPHERES).external icon
Updated 2021.
Hands-on Example:
- Phylogenetic tree.external icon
- Metadata table. Module 3.4 Demo Meta Data [CSV, 1KB]external icon
Note: Please first convert ToolkitModule_3.4-demoData.csv into a tsv file, according to the instructions here.external icon
Take the Feedback Survey for Module 3.4
email_03Get Email Updates
More modules and materials will be added to this toolkit, so please check back for updates or subscribe to the mailing list.
Related Videos
Page last reviewed: September 3, 2021
Content source: Centers for Disease Control and Prevention