At a glance
Sequencing technologies have revolutionized our ability to decode the DNA of disease-causing bacteria and viruses. The information we learn allows public health professionals to detect outbreaks sooner, including many outbreaks that would previously have gone undetected.
Decoding the whole genome
For more than 20 years, public health laboratories have used a DNA fingerprinting technology called Pulsed-Field Gel Electrophoresis (PFGE) to detect and track foodborne illness. In recent years, a set of new technologies have revolutionized our ability to decode DNA. Whole genome sequencing (WGS) gives us a much more detailed DNA fingerprint than PFGE. In public health, WGS transformed how epidemiologists and laboratory scientists approach the detection and investigation of outbreaks. This allows public health agencies across the US to detect outbreaks sooner, including many outbreaks that would previously gone undetected.
To show how this works, let's look at one example:
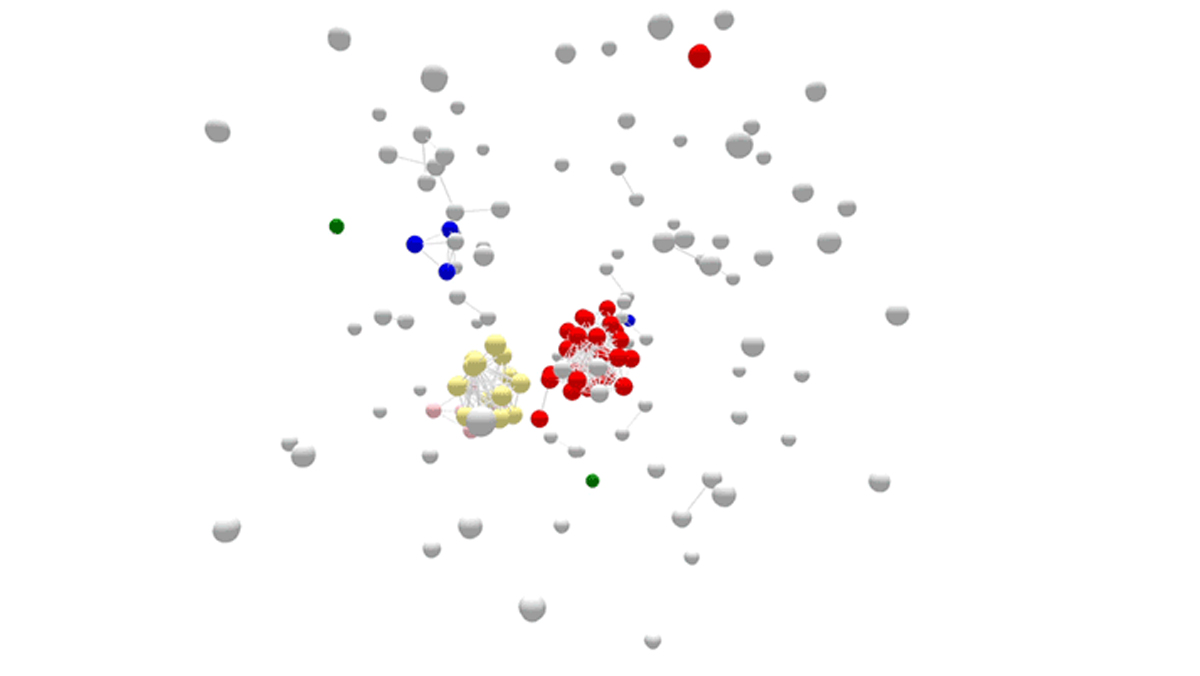
Unsorted cases
Over the course of year, several Salmonella outbreaks were identified, either in real time or retrospectively. Finding outbreaks is much easier and faster when related cases (shown in color) can be sorted from non-related cases (shown in grey).

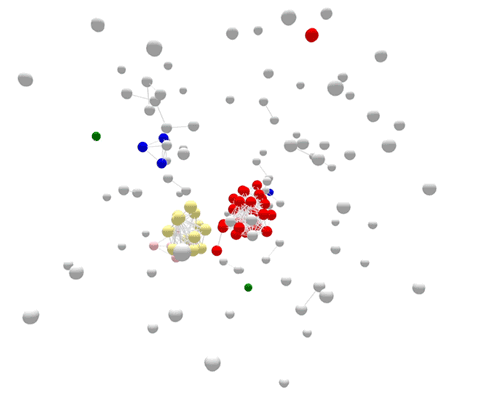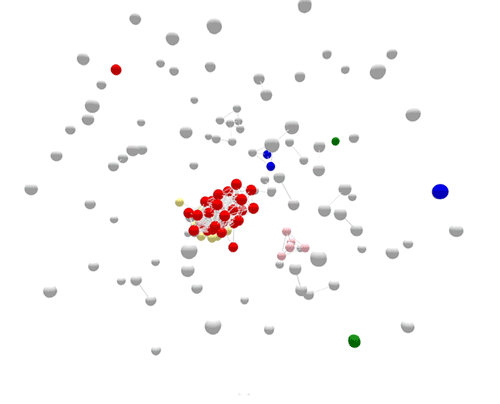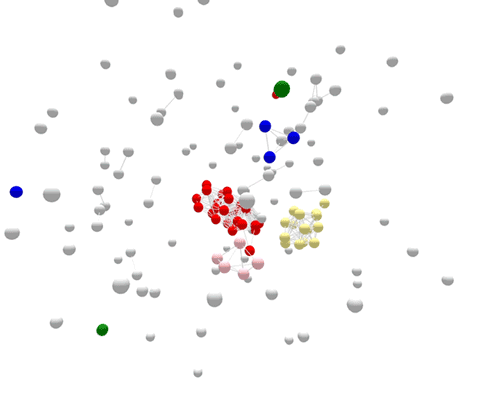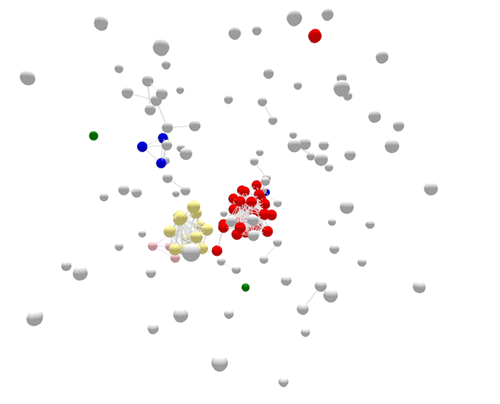
Cases sorted by PFGE
For the most part, PFGE accurately identifies related cases as are part of the same outbreak. However, each of the outbreak clusters include grey non-outbreak cases mixed in. Those unrelated cases complicate the investigation to find the common source.

Cases sorted by WGS
Using WGS, the outbreak cases are more tightly clustered, and stand out clearly from the disconnected, non-outbreak cases. The large (red) cluster in the center was initially connected to a small number of patrons at two different restaurants located in two separate states. Using WGS, investigators identified those cases were actually part of a larger outbreak, involving several patients who had not been to either restaurant.
The results
Epidemiologic data suggested the cause of the outbreak was shell eggs. Scientists used whole-genome sequencing to verify the source. Salmonella was found in the implicated eggs. The DNA fingerprint of the egg isolate matched the outbreak, confirming the attribution, and leading to a nationwide recall.
How WGS is being used
In the US, through the federally funded AMD Program, public health agencies apply next-generation sequencing in almost every area of infectious disease, such as:
In food safety, CDC works with FDA, USDA, NIH, and state and local public health agencies to quickly intervene in outbreaks and to better understand how to prevent pathogens from getting into the food system in the first place.
In flu, next-generation sequencing enables faster, more effective characterization of viruses to better understand how they emerge and to improve vaccine protection.
In viral hepatitis, next-generation sequencing is invaluable for outbreak investigations.
