COVID-19 Science Update released: March 5, 2021 Edition 79

The COVID-19 Science Update summarizes new and emerging scientific data for public health professionals to meet the challenges of this fast-moving pandemic. Weekly, staff from the CDC COVID-19 Response and the CDC Library systematically review literature in the WHO COVID-19 databaseexternal icon, and select publications and preprints for public health priority topics in the CDC Science Agenda for COVID-19 and CDC COVID-19 Response Health Equity Strategy.
Section headings in the COVID-19 Science Update align with the CDC Science Agenda for COVID-19.
Section headings in the COVID-19 Science Update have been changed to align
with the CDC Science Agenda for COVID-19.
Here you can find all previous COVID-19 Science Updates.
PEER-REVIEWED
Pregnancy and neonatal outcomes of COVID-19: co-reporting of common outcomes from PAN-COVID and AAP SONPM registries.external icon Mullins et al. Ultrasound in Obstetrics & Gynecology (February 23, 2021).
Key findings:
- Maternal deaths were uncommon in both the PAN-COVID (UK) and AAP SONPM (US) registries, 0.50% and 0.17%, respectively.
- Pre-term deliveries in the PAN-COVID cohort were higher than expected when compared with UK vital statistics records (12.0% vs. 7.5%) and higher in AAP SONPM when compared with 2018 US vital records (15.7% vs 10.0%).
- Neonatal deaths during the study time period were comparable to historical data.
Methods: Observational cohort study of pregnancy outcomes among women with SARS-CoV-2 infection using 2 national registries: the UK and Global Pregnancy and Neonatal outcomes in COVID-19 (PAN-COVID) registry including suspected and confirmed cases (n = 1,601), from January 1, 2020–July 25, 2020; and the US American Academy of Pediatrics Section on Neonatal Perinatal Medicine (AAP SONPM) National Perinatal COVID-19 registry (n = 2,399), from April 4, 2020–August 8, 2020. Limitations: Pre-term delivery rates may be inflated due to decisions to induce labor early to prevent adverse birth outcomes; limited inclusion of women with asymptomatic infection in the PAN-COVID study.
Implications: Pregnant women and women who plan on becoming pregnant should utilize all methods to reduce their risk of infection.
PREPRINTS (NOT PEER-REVIEWED)
Effectiveness of first dose of COVID-19 vaccines against hospital admissions in Scotland: national prospective cohort study of 5.4 million people.external icon Vasileiou et al. Lancet (February 19, 2021).
Key findings:
- Vaccines were protective against COVID-19 hospitalization; vaccine effect (VE) for BNT162b2 mRNA = 85% (95% CI 76%-91%); ChAdOx1 = 94% (95% CI 73%-99%).
- VE was observed over all time periods and increased over time, peaking 28–34 days post-vaccination for both the BNT162b2 mRNA (Pfizer-BioNTech) and ChAdOx1 (Oxford-AstraZeneca) vaccines.
- Similar findings observed in persons ≥80 years in an age-stratified pooled analysis for both vaccines.
Methods: A national, prospective cohort study using linked vaccination, testing, primary care, hospitalization, and mortality data for 5.4 million persons (99% of Scottish population). First doses of vaccine were administered between December 8, 2020–February 15, 2020 (1,137,775 persons). Data were analyzed with time dependent Cox and Poisson regression models fitted to estimate vaccine effectiveness against COVID-19 related hospitalization after the first dose. Limitations: Confounding due to differences within matched pairs; exclusion of certain groups like health care workers; other outcomes of interest not evaluated.
Implications: A single dose of Pfizer-BioNTech and Oxford-AstraZeneca vaccines resulted in substantial reductions in COVID-19-related hospitalizations in Scotland.
Here we present two reports (Zhou et al. and Supasa et al.) that examine the potential impacts of two recently emerged and regionally dominant SARS-CoV-2 variants (B.1.351 and B.1.1.7) on the effectiveness of existing prevention and treatment options for COVID-19 infection.
PEER-REVIEWED
A. Evidence of escape of SARS-CoV-2 variant B.1.351 from natural and vaccine induced sera.external icon Zhou et al. Cell (February 23, 2021).
Key findings:
- With convalescent plasma, neutralization titers against B.1.351 were ~13.3-fold reduced compared to wild-type.
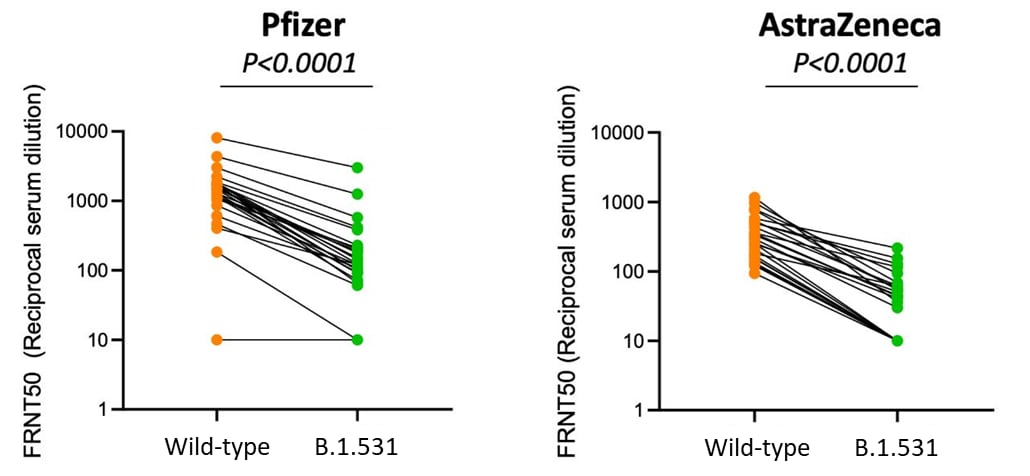
- With sera from Pfizer-BioNTech and Oxford-AstraZeneca vaccinees, geometric mean titers for B.1.351 were 7.6-fold and 9-fold lower compared to a wild-type SARS-CoV-2, respectively (Figure).
- 14/20 monoclonal antibodies (mAbs) had >10-fold fall in neutralization titers; most showed a complete knock out of activity.
- Neutralization of REGN10933 (now available under EUA as casirivimab given in combination with imdevimabexternal icon) was severely impaired 773-fold against B.1.351.
Figure:
Note: Adapted from Zhou et al. Comparison of FRNT50 titers between B.1.531 variant and wild-type SARS-CoV-2 for the Pfizer-BionNTech and Oxford-AstraZeneca vaccines. Higher FRNT50 indicate better neutralization of virus. Licensed under CC BY.
B. Reduced neutralization of SARS-CoV-2 B.1.1.7 variant by convalescent and vaccine seraexternal icon. Supasa et al. Cell (February 18, 2021).
Key findings:
- An amino acid substitution (N501Y) in the spike protein of SARS-CoV-2 variant B.1.1.7 caused numerous functional changes:
- 7-fold increased binding to human ACE2 receptors.
- Decreased viral neutralization by numerous antibodies.
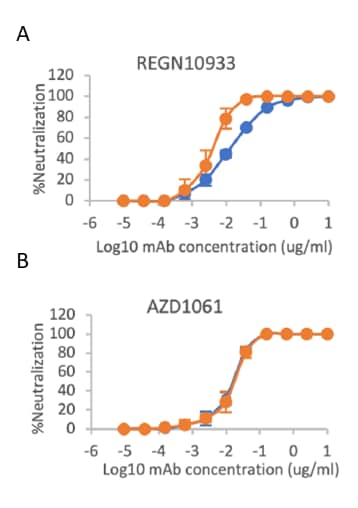
- Widespread escape of viral neutralization was not observed for commercially available monoclonal antibodies (Figure), or for sera from vaccinees or from convalescent patients.
Figure:
Note: Adapted from Supasa et al. Percent neutralization of wild-type SARS-CoV-2 or B.1.1.7 variant vs. commercially available monoclonal antibody concentration (mAb) (A. Regeneron: REGN10933 and B. AstraZeneca: AZD106). Licensed under CC BY.
a
Methods for both studies (Zhou et al. and Supasa et al.): In two separate studies, the neutralization of B.1.351 and of B.1.1.7 along with a wild-type SARS-CoV-2 viruses (control) were compared in vitro using a focus reduction neutralization test (FRNT) with immune sera and plasma from a UK cohort of persons recovered from or vaccinated against SARS-CoV-2 as well as monoclonal antibody (mAbs) cocktails. Additionally, for the B.1.1.7 variant, the N501Y mutation was modeled to examine effects on antibody and ACE2 affinity. Limitations for both studies: In vitro neutralization assays do not include assessment of the contributions to in vivo protection provide by other parts of the immune response such as B or T cells. Samples used in the neutralization studies were taken early after illness or vaccination and protective levels may change over time.
Implications for both studies (Zhou et al. and Supasa et al.): In Zhou et alexternal icon., neutralizing activity of antibodies were lower against SARS-CoV-2 B.1.351 than wild-type SARS-CoV-2 viruses suggesting monoclonal antibody therapy and vaccine effectiveness will be impacted. In Supasa et al.external icon, the B.1.1.7 variant did not evade treatment with existing monoclonal antibodies and there was no evidence of vaccine escape, indicating protection from the B.1.1.7 variant by existing vaccines. Wu et al.external icon also found evidence that the Moderna vaccine-induced antibody response neutralized the B.1.1.7 variant in vitro but less so the B.1.351 variant.
PREPRINTS (NOT PEER-REVIEWED)
Acquisition of the L452R mutation in the ACE2-binding interface of Spike protein triggers recent massive expansion of SARS-Cov-2 variants.external icon Tchesnokova et al. bioRxiv (February 22, 2021).
Key findings:
- L452R is associated with a CAL.20C viral variant (lineage B.1.429), that since November-December 2020 has been associated with multiple outbreaks and spread across California (Figure).
- A L452R-carrying CAL.20A variant (lineage B.1.232) emerged more recently and is also circulating primarily in California.
- Several independent L452R-carrying lineages have recently emerged internationally; >90% of these isolates were reported between December 2020–February 2021.
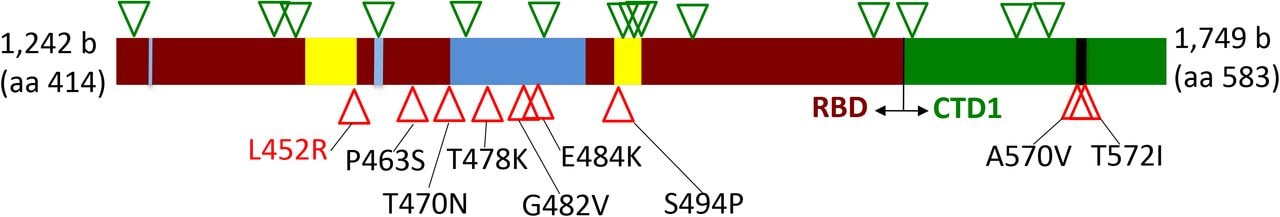
Methods: Sequenced the spike protein 414-583 amino acid region (Figure), partially encompassing the ACE2 receptor-binding domain of the SARS-CoV-2 spike protein, from a subset of 570 SARS-CoV-2 RNA samples isolated between April 2020 and February 2021, from Washington, California, Arizona, Colorado, Minnesota and Illinois. Sequences from the Global Initiative on Sharing Avian Influenza Data (GISAIDexternal icon) database were checked for L452 substitutions. Limitations: Possible sample collection bias; relatively small sample size; limited to publicly available genomes.
Implications: Emergence of the L452R mutation suggests potential adaptive value to SARS-CoV-2. Further research is needed to determine whether L452R facilitates escape from neutralizing antibodies and increased infectivity.
Figure:
Note: Adapted from Tchesnokova et al. Triangles show distribution of silent mutations (i.e., base change did not result in amino acid change) and amino acid mutations (i.e., base change led to change in amino acid) across regions 414-583 of the spike protein. Bar shows the receptor-binding domain (RBD), the C-terminal domain of S1 spike region, the receptor-binding ridge epitope residues, the 443-450 loop residues, and the 570-572 loop residues. Used by permission of authors.
Several SARS-CoV-2 potential variants of interest have been identified recently. Two reports (West et al. and Annavajhala et al.) describe emergence of the B.1.526 variant in New York and the northeastern United States.
A. SARS-CoV-2 lineage B.1.526 emerging in the New York region detected by software utility created to query the spike mutational landscape.external icon West et al. bioRxiv (February 23, 2021).
Key findings:
- In New York state during November 2020-February 2021, B.1.526 isolates (with spike mutations T95I and D253G) increased from 0.2% to 27.8% of sequenced SARS-CoV-2 isolates.
- In New York state during December 2020–February 2021, B.1.526 isolates (with the spike mutations L5F, T95I, D253G, E484K, D614G, and A701V) increased from 0.9% to 13.9% of sequenced SARS-CoV-2 isolates.
Methods: Developed a software tool, Variant Database, to detect changes in the spike protein mutational landscape by identifying collections of isolate sequences (clusters) reported to the Global Initiative on Sharing Avian Influenza Data (GISAIDexternal icon). Limitations: Only a small proportion of SARS-CoV-2 infections are tested for genetic sequences and reported to GISAID; geographic sampling might have varied over time and was not necessarily random (i.e., representative).
B. A novel SARS-CoV-2 variant of concern, B.1.526, identified in New York.external icon Annavajhala et al. medRxiv (February 25, 2021).
Key findings:
- A novel variant, B.1.526, has been identified in New York. In a subset of samples flagged as potential N501Y- or E484K-harboring strains, 75% of the E484K isolates fell within the B.1.526 lineage.
- Nearly all B.1.526 variants identified had common mutations in the spike protein (L5F, T95I, D253G, E484K, D614G, and A710V).
- ~140 genomes, highly related to the B.1.526 variant, were found in public databases; sampled predominantly from the northeastern US.
Methods: A random sample of SARS-CoV-2 nasopharyngeal swab specimens (n = 1,142) from 4,358 infected patients at a quaternary care center in New York City during November 1, 2020–February 15, 2021 were assayed for N501Y and E484K mutations. Whole genome nanopore sequencing was done on those flagged as potential N501Y- or E484K-harboring strains (n = 65), along with samples negative for these mutational signatures with Ct values <35 (n = 65). Limitations: Single center; not geographically generalizable; subset of positive samples were tested for the two mutations; study based on clinical submissions.
Implications for both studies (West et al. and Annavajhala et al.): The B.1.526 variant emerged rapidly and may already be widespread in the northeastern US. This variant includes multiple mutations that might facilitate SARS-CoV-2 infection and spread, including through attenuation of antibody neutralization (E484K), escape from antibodies against the N-terminal domain (D253G), and ACE2 interactions that might enhance viral infectivity (S477N). Continued mitigation efforts and increased capacity for genetic surveillance will be critical to limit and detect further development and spread of mutations.
PEER-REVIEWED
BNT162b2 mRNA Covid-19 Vaccine in a Nationwide Mass Vaccination Settingexternal icon. Dagan et al. NEJM (February 24, 2021).
Key findings:
- 14-20 days after first dose and 7 or more days after second dose, respectively, BNT162b2 vaccination prevented an estimated:
- 46% and 92% of documented infection.
- 57% and 94% of symptomatic Infection.
- 74% and 87% of hospitalization.
- 62% and 92% of severe Disease.
- 72% of deaths (after first dose) from COVID-19 (figure).
Methods: Observational matched cohort study of mass vaccination campaign in Israel used data from the largest health care organization (53% of population) to determine COVID-19 vaccine (BNT162b2) effectiveness on 5 outcomes. All persons newly vaccinated from December 20–February 1, 2021 (n = 596,618) were matched 1:1 to unvaccinated controls (n = 596,618). Limitations: Confounding due to differences between unvaccinated controls and vaccinated persons; exclusion of certain groups like healthcare workers; adverse advents not addressed.
Implications: These data provide evidence of real-world effectiveness of the BNT162b2 vaccine for a wide range of COVID-19 outcomes including severe outcomes and death, suggesting that widespread vaccination can mitigate COVID-19 burden.
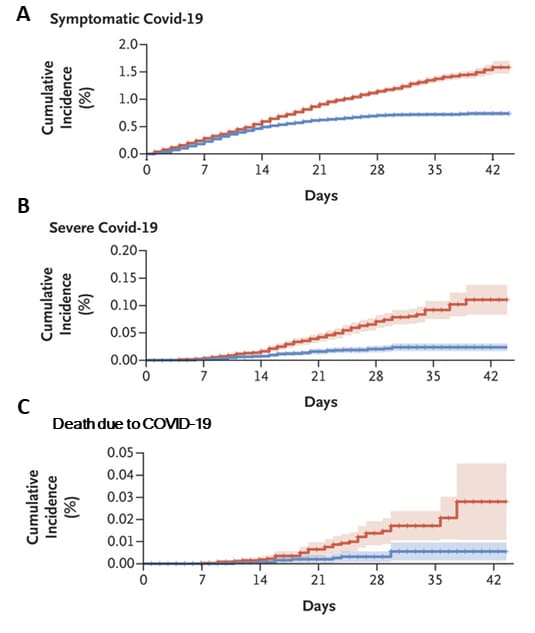
Figure:
Note: Adapted from Dagan et al. Cumulative incidence curves for COVID-19 outcomes (A. Symptomatic COVID-19, B. Severe COVID-19, C. Death due to COVID-19) among vaccinated and unvaccinated persons, starting from the first dose of vaccination. Shaded areas represent 95% confidence intervals. From NEJM, Dagan et al., BNT162b2 mRNA Covid-19 Vaccine in a Nationwide Mass Vaccination Setting. Copyright© (2021) Massachusetts Medical Society. Reprinted with permission from Massachusetts Medical Society.
Association of the timing of school closings and behavioral changes with the evolution of the coronavirus disease 2019 pandemic in the US.external icon Zimmerman et al. JAMA Pediatrics (February 22, 2021).
Key findings:
- A 2-week delay in school closure alone was associated with an additional 587,000 cases (95% CI 213,000-1,153,000) and 23,000 deaths (95% CI 2,000-62,000).
- a 2-week delay in voluntary behavior change was associated with an additional 4,300,000 cases (95% CI 2,542,000-6,995,000) and 140,000 deaths (95% CI 65,000-294,000) (Figure).
Methods: A population-based interrupted-time-series analysis to simulate the independent association of school closures and behavioral changes with state-level COVID-19 case and death data (New York Times) from March 8–May 18, 2020. Proxies of behavioral change were time at work, time at home, searches for hand sanitizer, and dining out at restaurants based on cell phone and internet data. Limitations: Data for restaurant/dining out was only available for 36 states; methods of data collection could lead to measurement error for behaviors; did not account for differences in county-level school closure. Editorial Note: There are a few instances in the paper where results are reported as “per each day of delay in school closure, there was an associated reduction in cases”. However, the author confirmed that the correct language is “for each day earlier a school was closed, there was an associated reduction in cases.”
Implications: Voluntary behavioral changes such as reductions in time spent at work have a greater impact on COVID-19 spread than school closures which modestly reduce COVID-19 incidence and mortality.
Figure:
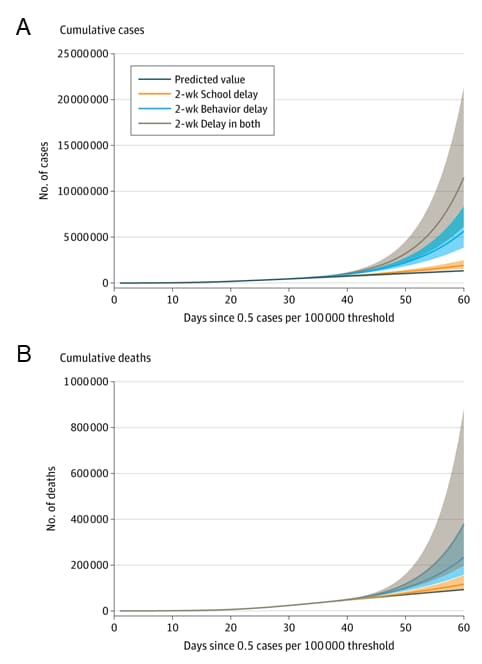
Note: Adapted from Zimmerman et al. Modeled coronavirus disease 2019 (A) cases and (B) death under various scenarios. Scenarios included: Predicted value (baseline estimated value), 2-week delay in school closures, 2-week delay in behavioral changes, and a 2-week delay in both school closures and behavioral changes since cases were 0.5 per 100,000. Shading indicates 95% CIs derived from Monte Carlo simulation with 1,000 trials. Reproduced with permission from JAMA Pediatrics. 2021. doi:10.1001/jamapediatrics.2020.6371. Copyright© (2021) American Medical Association. All rights reserved.
Detection, Burden, and Impact
- Barak et al. Lessons from applied large-scale pooling of 133,816 SARS-CoV-2 RT-PCR testsexternal icon. Science Translational Medicine (February 22, 2021). SARS-CoV-2 samples were tested in pools between March and mid-September 2020, sparing 76% of individual RNA extraction and RT-PCR tests, demonstrating strong support for the use of pooling in high throughput SARS-CoV-2 testing.
- Yaren et al. Ultra-rapid detection of SARS-CoV-2 in public workspace environments.external icon PLoS One (February 24, 2021). A rapid test for entry into schools and workspaces based on a displaceable probe loop amplification architecture is described. This test trades sensitivity, that may not be necessary for screening, for speed, requires minimal sample manipulation, can be performed by a layperson person and is inexpensive.
- Vandergaastrgaast et al. IMMUNO-COV™ v.2.0: development and validation of high-throughput clinical assay for measuring SARS-CoV-2 neutralizing antibody titers.external icon medRxiv (Preprint, February 19, 2021). IMMUNO-COV v.2.0 was developed and validated as a scalable, high-throughput assay that makes clinical determination of neutralizing antibodies feasible.
- Kojaku et al. The effectiveness of backward contact tracing in networksexternal icon. Nature Physics (February 25, 2021). ‘Backward’ contact tracing is more effective than ‘forward’ tracing because it leverages social network bias; it is exceptionally effective for identifying super-spreader individuals and events.
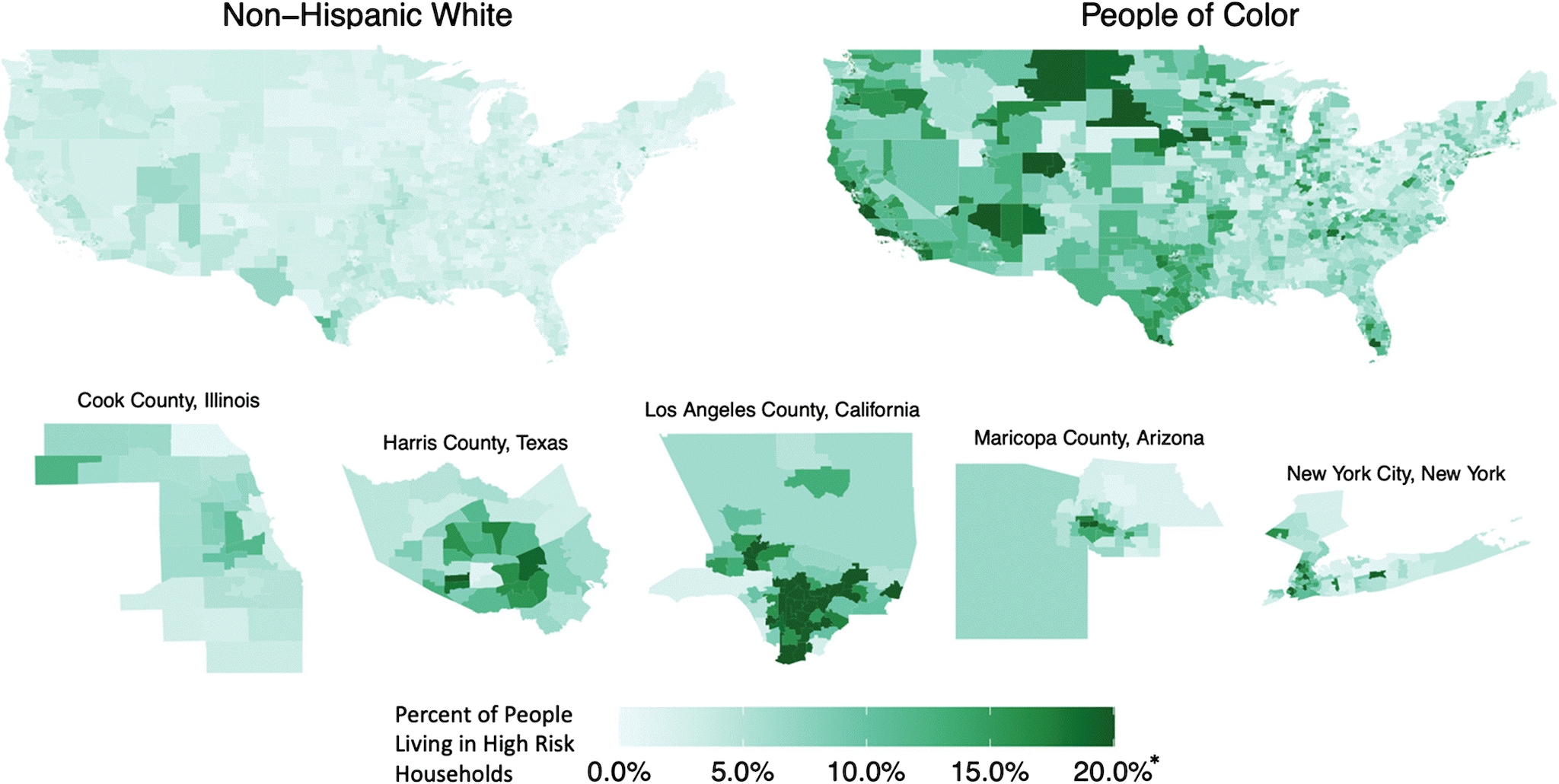
- Reitsma et al. Mapping inequality in SARS-CoV-2 household exposure and transmission risk in the USAexternal icon. Journal of General Internal Medicine (February 18, 2021). Maps of structural inequality were created using US Census Data to identify high-risk households (with an essential worker, more people than rooms, multigenerational) which are found disproportionately in communities of color.
Figure:
Note: Adapted from Reitsma et al. Percent of people living with at least one essential worker in a household with fewer rooms than people, by race/ethnicity. Inset shows estimates for the four most-populated counties and New York City. Permission request in process.
Transmission of SARS-CoV-2
- Kraay et al. Risk for fomite-mediated transmission of SARS-CoV-2 in child daycares, schools, nursing homes, and offices. Emerging Infectious Diseases (February 24, 2021). SARS-CoV-2 virus can persist on surfaces, and fomites might be a substantial source of transmission risk – especially in schools and in daycare settings. Per this modeling study, hourly cleaning and disinfection may interrupt fomite transmission in office settings but may not be adequate in child daycares and schools.
Natural History of SARS-CoV-2 Infection
- Challen et al. Increased hazard of mortality in cases compatible with SARS-CoV-2 variant of concern 202012/1 – a matched cohort study.external icon medRxiv (Preprint, March 1, 2021). The SARS-CoV-2 lineage B.1.1.7 (identified by Public Health England as VOC-202012/1) is associated with a higher risk of mortality (hazard ratio 1.7 [95% CI 1.3-2.2]) compared to the wild-type lineage.
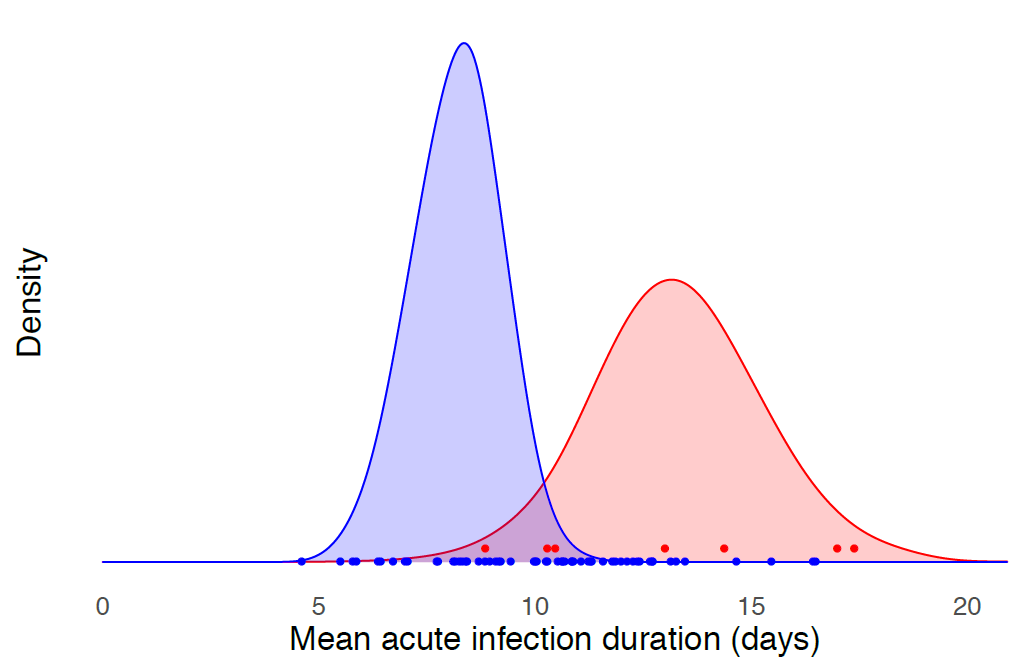
- Kissler et al. Densely sampled viral trajectories suggest longer duration of acute infection with B.1.1.7 variant relative to non-B.1.1.7 SARS-CoV2.external icon medRxiv (Preprint, February 19, 2021). A longitudinal assessment of SARS-CoV-2 viral loads found that SARS-CoV-2 variant B.1.1.7 may cause longer acute phase infections with similar peak viral concentrations as non-B.1.1.7 SARS-CoV-2. The extended duration may contribute to increased transmissibility.
Figure:
Note: Adapted from Kissler et al. Estimated viral trajectories for B.1.1.7 and non-B.1.1.7 SARS-CoV-2. Mean acute infection duration (days). Licensed under CC BY-NC-ND 4.0.
- Karbiener et al. Longitudinal analysis of SARS-CoV-2 antibodies in 8000 U.S. first-time convalescent plasma donations.external icon Transfusion (February 22, 2021). From April through September 2020, there was a marked decrease in average SARS-CoV-2 IgG values in COVID-19 convalescent plasma from first-time US plasma donors due to increases in donations from progressively younger donors who tend to have less severe COVID-19.
Prevention, Mitigation, and Intervention Strategies
- Hall et al. Effectiveness of BNT162b2 mRNA vaccine against infection and COVID-19 vaccine coverage in healthcare workers in England, multicentre prospective cohort study (the SIREN study).external icon Lancet (Preprint, February 22, 2021). With B.1.1.7 as the dominant variant in circulation, the BNT162b2 mRNA vaccine effectively prevented both symptomatic and asymptomatic infection in working age adults in a cohort of healthcare workers.
- Ebinger et al. Prior COVID-19 infection and antibody response to single versus double dose mRNA SARS-CoV-2 vaccination.external icon medRxiv (Preprint, February 24, 2021). Vaccine (Pifzer-BioNTech mRNA) antibody response evaluated in a large cohort of healthcare workers with and without prior infection provides evidence that a single dose of mRNA vaccine could provide immunity in previously infected persons comparable to that seen in naïve persons following a second dose.
- Nelson et al. SARS-CoV-2 positivity on or after 9 days among quarantined student contacts of confirmed cases.external icon JAMA (February 19, 2021). Compared with a 14-day quarantine without testing, the 9-day quarantine with testing protocol instituted for student contacts of confirmed cases resulted in earlier return to school without risk for subsequent illness.
Disclaimer: The purpose of the CDC COVID-19 Science Update is to share public health articles with public health agencies and departments for informational and educational purposes. Materials listed in this Science Update are selected to provide awareness of relevant public health literature. A material’s inclusion and the material itself provided here in full or in part, does not necessarily represent the views of the U.S. Department of Health and Human Services or the CDC, nor does it necessarily imply endorsement of methods or findings. While much of the COVID-19 literature is open access or otherwise freely available, it is the responsibility of the third-party user to determine whether any intellectual property rights govern the use of materials in this Science Update prior to use or distribution. Findings are based on research available at the time of this publication and may be subject to change.