Frequently Asked Questions
What is PulseNet?
PulseNet is a national laboratory network consisting of 83 public health and food regulatory laboratories. PulseNet groups together people who most likely ate the same contaminated food, or who were exposed to illness-causing microorganisms in some other way. The network does this by analyzing DNA fingerprinting on the bacteria making people sick, and on the bacteria found in food and the environment. People and foods infected or contaminated with bacteria with the same fingerprint are likely to be part of an outbreak. Public health and regulatory investigators use this information to identify the source of illness, such as an unrecognized problem in the food supply chain. Microbiologists generate these fingerprints using molecular subtyping tools such as pulsed-field gel electrophoresis (PFGE), multiple-locus variable number tandem repeat analysis (MLVA), and whole genome sequencing (WGS). Associated networks, including CaliciNet and CryptoNet have similar functions for norovirus and Cryptosporidium parasites.
Every state has at least one public health laboratoryexternal icon that can match up bacteria from sick people in many different locations using PulseNet’s DNA fingerprinting techniques and databases. PulseNet tracks what is being reported to CDC in real time, compares it to what was reported in the past, and looks for any increases that could signal an outbreak. The PulseNet databases keep growing for this reason. They contain over half a million fingerprints from human illness, food, and the environment.
- PulseNet has revolutionized how we detect and investigate foodborne disease outbreaks since it started in 1996. Before PulseNet, these outbreaks often went undetected or were discovered only after they grew very large.
- PulseNet is undergoing a major technological transformation from 20 years of PFGE data to a more powerful and precise method, WGS. This will improve the network’s ability to find and investigate outbreaks. Implementing whole genome sequencing will provide the most accurate bacterial fingerprinting data possible today.
What is PulseNet’s role?

PulseNet identifies clusters of illness, or groups of persons ill with the identical bacteria that may be the cause of an outbreak. Once a cluster is identified, PulseNet works with laboratory and epidemiological investigations teams at CDC, the federal food regulatory agencies – the U.S. Food and Drug Administration (FDAexternal icon) and the U.S. Department of Agriculture’s Food Safety and Inspection Service (USDA/FSISexternal icon) – and state and local health departments to figure out what is making people sick. Each year PulseNet identifies:
- About 1,500 clusters of foodborne disease at local or state levels
- About 250 clusters that span multiple states
- About 30 multistate outbreaks that are linked to a food source
PulseNet detects outbreaks rapidly making it possible to recall and remove thousands of contaminated food products, totaling over a billion pounds. Most importantly, many products and services are safer today because of investigations PulseNet initiated. These outbreak investigations have stimulated production changes in the food industry and helped federal agencies create new or improved guidance, policy, and regulations that have prevented thousands of foodborne illnesses making our food supply safer.
How do we detect clusters?
The graph below shows groups of matching patterns, or clusters, of a common serotype of Salmonella over a year in the United States. PulseNet database managers create these graphs and look for clusters that are above the expected number of patterns seen in a month. In this example the database manager would investigate and notify an epidemiologist in the month of December that this cluster could be an outbreak.
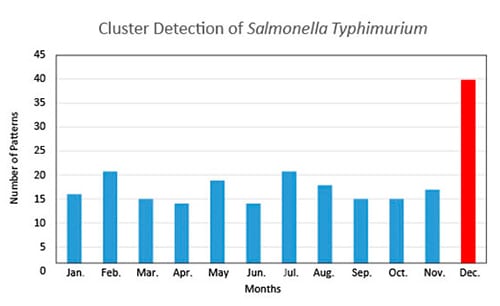
Why was PulseNet developed?
In 1993, more than 700 people fell ill and four died in the western United States after eating at restaurants of a large, regional fast food chain. More than a month after the first person fell ill, investigators found the cause—a bacterium called Escherichia coli O157:H7. Two weeks later, burger patties were identified as the culprit. During this outbreak, CDC scientists performed PFGE to determine that the strain of E. coli O157:H7 found in patients was the same as the strain found in the hamburgers.
CDC scientists decided that outbreaks could be identified and stopped sooner if all public health laboratories could perform the same DNA fingerprinting tests on bacteria from patients and share results with all other laboratories. This idea was the beginnings of PulseNet.
CDC worked with the Association of Public Health Laboratoriesexternal icon(APHL) to develop PulseNet. The goal was to get a DNA fingerprint of all foodborne bacteria as they were submitted to public health laboratories in the states, gather the information locally and nationally, and continuously monitor these databases for clusters of cases with the same DNA fingerprints to speed up the recognition and investigation of outbreaks.
PulseNet achieved this by implementing the same methods for testing bacteria at all participating laboratories which ensured that the results, or fingerprints, could be compared in the shared databases.

Why is PulseNet important to the public’s health?
- PulseNet helps public health scientists determine where our food safety systems have failed so that solutions can be developed to prevent future illnesses.
- PulseNet also helps scientists determine whether an outbreak is occurring, even if only a few people are ill or if they are geographically far apart.
- The network allows us to identify outbreaks and their causes in a matter of hours rather than days or weeks.
- Epidemiologists across the country use PulseNet data to help determine what specifically is making people sick. They also work with public health regulatory agencies, such as the U.S. Food and Drug Administration (FDAexternal icon) and the U.S. Department of Agriculture’s Food Safety and Inspection Service (USDA/FSISexternal icon), to find solutions to end outbreaks and prevent them from ever happening. These solutions include immediate measures, such as food recalls, and long-term measures, such as enhanced guidance, policies, and regulations that lead to new production practice.
Why is PulseNet effective?
PulseNet is effective because all of the laboratories in its network subtype all foodborne bacteria in real time and follow the same procedures using the same standards. As a result, we can compare fingerprints generated by different laboratories. In PulseNet, the quality and uniformity of the data are ensured by a quality assurance and quality control program.
How does PulseNet work?
- A person falls ill and visits a doctor.
- The doctor suspects a foodborne illness and asks for a sample (usually a stool sample), which he or she sends to a laboratory at a clinic or hospital.
- The clinical laboratory processes the sample to isolate (pull out) the bacteria that is making the patient ill—for example, Salmonella.
- The clinical laboratory notifies the doctor, who will tell the patient and discuss treatment options. The clinical laboratory also sends the sample to a local or state public health laboratory.
- The public health laboratory determines what kind of Salmonella it is—for example, Salmonella serotype Enteritidis. Public health laboratoriesexternal icon are a first line of defense to protect the public against foodborne illnesses and other health hazards.
- The public health laboratory produces a DNA fingerprint of the bacteria, traditionally using a process called pulsed-field gel electrophoresis (PFGE), to get the unique pattern.
- The public health laboratory uploads the pattern into an electronic database in its laboratory and also to the national databases at CDC.
- Microbiologists and epidemiologists review the laboratory reports to decide whether something unusual or unexpected is occurring that warrants further investigation.
- Epidemiologists interview the patients. These interviews may gather simple facts of their illness, or they may ask about everything they ate before they got sick, where they have been, and other questions that might provide a clue as to what caused their illness.
- The microbiologists continuously search the databases for identical patterns. For example, if they find a pattern—which comes from five different samples, from five different ill people—that appears to be the same, it is a clue that these five people may be part of a single outbreak, caused by a single food (or other source).
- These groups of matching patterns, called clusters, spur investigations by local, state, and national agencies to identify the source of the outbreak. To identify specific causes of the outbreak, they can:
- Compare answers to interview questions among ill people in the same cluster.
- Connect PFGE patterns in patients to patterns found in food monitoring programs.
- Conduct epidemiological studies.
- When these agencies are able to identify an outbreak’s source, they can collaborate to control further spread of the outbreak and to develop strategies to prevent the outbreak from happening again.
How does DNA fingerprinting help outbreak investigations?
What we eat and how we eat in the United States has changed. In the early 1900s, most food was consumed close to where it was produced. Food safety gaps were usually discovered only when groups of people in the same location became ill, such as picnics or schools. Then and now, such outbreaks are caused by unsafe food preparation practices, such as improper cooking or refrigeration. In the last half-century, food production has become increasingly centralized, and food products are often transported great distances before arriving at our dinner tables. Illnesses caused by errors in food production can sicken people over a wide area and may not be recognized as a problem in any one community. PulseNet is helping to change that. Since PulseNet began 20 years ago, we have seen a dramatic increase in our ability to detect widespread outbreaks that occur across many communities, even when there are only a few illnesses in each community.
PulseNet strengthens our ability to identify and investigate outbreaks by
- Identifying which illnesses are truly part of an outbreak. DNA fingerprinting combined with epidemiology information can help differentiate outbreak-associated illnesses from unrelated illnesses occurring at the same time.
- Detecting national outbreaks through surveillance. DNA fingerprinting can link bacteria found in sick people in one community to bacteria found in sick people around the country. Those fingerprints connect the dots to help find outbreaks.
Isn’t it better to prevent outbreaks before they happen?
Preventing outbreaks from occurring is the ultimate goal of food safety policies and programs, including good manufacturing practices, food safety inspections, and hazard analysis and critical control pointsexternal icon plans. Because of the complexity of food production, distribution, and preparation, we may not be able to completely eliminate foodborne disease, but PulseNet will allow us to quickly stop outbreaks and save lives.
PulseNet, with other foodborne illness surveillance systems, is our best tool for detecting problems within our food safety system that otherwise would go unrecognized. Identifying these problems allows us to take rapid action in the short term, to identify the contaminated food and remove it from the market; in the longer term, by helping pinpoint defective food manufacturing practices which can be altered, thereby rendering the food supply safer.