Outbreaks of Highly Pathogenic Avian Influenza (H5N6) Virus Subclade 2.3.4.4h in Swans, Xinjiang, Western China, 2020
Yanbing Li, Minghui Li, Yulei Li, Jingman Tian, Xiaoli Bai, Cen Yang, Jianzhong Shi, Ridengcaicike Ai, Weidong Chen, Wentao Zhang, Jie Li, Yufei Kong, Yuntao Guan, and Hualan Chen
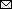
Author affiliations: State Key Laboratory of Veterinary Biotechnology, Harbin Veterinary Research Institute, Chinese Academy of Agricultural Sciences, Harbin, Heilongjiang, China (Y. Li, M. Li, Y. Li, J. Tian, X. Bai, J. Shi, C. Yang, Y. Kong, Y. Guan, H. Chen); Preventive and Control Center for Animal Disease of Xinjiang Uyghur Autonomous Region, Urumqi, Xinjiang Wei Autonomous Region, China (R. Ai, J. Li); Preventive and Control Center for Animal Disease of Xinjiang Crops, Urumqi (W. Zhang, J. Li)
Main Article
Figure 1

Figure 1. Geography and phylogeny of avian influenza (H5N6) outbreaks among migratory whooper swans (Cygnus cygnus) and mute swans (C. olor), Xinjiang Province, China, January 2020. A) Disease outbreak sites are marked with red drops, and dates of the outbreaks are indicated. Inset map shows islands in the South China Sea. B) Phylogenetic tree of the hemagglutinin (HA) genes of H5 viruses. The HA gene maximum clade credibility tree of the H5 viruses was constructed by using the BEAST 1.8.4 software package (https://beast-dev.github.io/beast-mcmc). Node bars indicate 95% highest posterior density of the node height. Each branch is colored by posterior probability: the 13 H5N6 viruses reported in this study are shown in red and the HA donor of the H5N1 vaccine Re-11 in green. The time to the most recent common ancestor is labeled at the bottom of the tree, which was estimated by using the Bayesian Markov chain Monte Carlo method in the BEAST 1.8.4 software package.
Main Article
Page created: September 02, 2020
Page updated: November 19, 2020
Page reviewed: November 19, 2020
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.
