 |
|
|
|
|
|
|
| ||||||||||
|
|
|
|
|
|
|
||||
| ||||||||||
|
|
|
|
|
Persons using assistive technology might not be able to fully access information in this file. For assistance, please send e-mail to: mmwrq@cdc.gov. Type 508 Accommodation and the title of the report in the subject line of e-mail. Removing a Barrier to Computer-Based Outbreak and Disease Surveillance --- The RODS Open Source ProjectJeremy U. Espino, M. Wagner, C. Szczepaniak, F-C. Tsui, H. Su, R. Olszewski, Z. Liu, W. Chapman, X. Zeng, L. Ma, Z. Lu, J. Dara
Corresponding author: Jeremy U. Espino, Real-Time Outbreak and Disease Surveillance Laboratory, University of Pittsburgh, Suite 500, Cellomics Building, 500 Technology Dr., Pittsburgh, PA 15219. Telephone: 412-383-8130; Fax: 412-383-8135; E-mail: jue@cbmi.pitt.edu. AbstractIntroduction: Computer-based outbreak and disease surveillance requires high-quality software that is well-supported and affordable. Developing software in an open-source framework, which entails free distribution and use of software and continuous, community-based software development, can produce software with such characteristics, and can do so rapidly. Objectives: The objective of the Real-Time Outbreak and Disease Surveillance (RODS) Open Source Project is to accelerate the deployment of computer-based outbreak and disease surveillance systems by writing software and catalyzing the formation of a community of users, developers, consultants, and scientists who support its use. Methods: The University of Pittsburgh seeded the Open Source Project by releasing the RODS software under the GNU General Public License. An infrastructure was created, consisting of a website, mailing lists for developers and users, designated software developers, and shared code-development tools. These resources are intended to encourage growth of the Open Source Project community. Progress is measured by assessing website usage, number of software downloads, number of inquiries, number of system deployments, and number of new features or modules added to the code base. Results: During September--November 2003, users generated 5,370 page views of the project website, 59 software downloads, 20 inquiries, one new deployment, and addition of four features. Conclusions: Thus far, health departments and companies have been more interested in using the software as is than in customizing or developing new features. The RODS laboratory anticipates that after initial installation has been completed, health departments and companies will begin to customize the software and contribute their enhancements to the public code base. IntroductionIn October 1999, researchers at the University of Pittsburgh began developing the Real-Time Outbreak and Disease Surveillance system (RODS), with the goal of improving public health agencies' capability to detect a specific threat: a large-scale, surreptitious release of Bacillus anthracis. The rate of this technology's adoption, although accelerating, is not commensurate with the severity of the health threats posed by biologic terrorism, emerging infections, and common disease outbreaks. Such threats warrant rapid deployment; therefore, barriers to the technology's adoption need to be identified and removed. This paper describes the evolution of the RODS system, previous efforts to transition the technology, and the rationale behind the creation of an open-source project. It also describes how the software is licensed, the infrastructure created to enable growth of the RODS open-source community, efforts to publicize the project, metrics collected to assess its progress, the software architecture of the latest version of RODS, and plans for additional software development. RODS System DescriptionThe first version of RODS collected patient chief-complaint data from eight hospitals in a single health-care system via Health Level 7 (HL7) (1) messages in real time, categorized these data into syndrome categories by using a classifier based on International Classification of Diseases, Ninth Revision (ICD-9) codes, aggregated the data into daily syndrome counts, and analyzed the data for anomalies possibly indicative of disease outbreaks. The system provided an Internet-based interface enabling users to view the data in graphs and maps (Figure 1). After demonstrating the feasibility of such a system within a single health-care system in Pittsburgh and conducting research to support the hypothesis that such a system could detect disease outbreaks (2,3), RODS' developers expanded the system to collect additional data types and then deployed RODS in multiple states. The application service provider (ASP) version of RODS at the University of Pittsburgh collects de-identified chief complaints from 76 hospitals in Pennsylvania, Utah, and Ohio (4,5) and also serves as the user interface for the National Retail Data Monitor (NRDM), which collects and analyzes daily sales data for over-the-counter (OTC) medication sales (6,7). The feasibility of rapid deployment of RODS was demonstrated during the 2002 Winter Olympics in Salt Lake City, Utah (4,8,9). In addition, the capability to integrate other surveillance data types (e.g., electronic laboratory reports [10], free-text chief complaints (11,12), laboratory orders, dictated radiology reports, dictated hospital reports [13--15], and poison control center calls [16]) was added. Much of the code (originally in Perl and C) was rewritten in Java,™ and basic research was conducted on data and algorithms relevant to this emerging science (17). Technology TransitionThe initial effort to make RODS software available involved licensing it for noncommercial use. In December 2002, the University of Pittsburgh began offering the RODS system as compiled byte code, free of charge to public health departments. To date, >180 downloads of this version of the RODS system and >200 downloads of the Bayesian parser have been counted. Despite reports of successful installations in Hong Kong [David Wong, Hong Kong RODS Team, personal communication, May 15, 2003] and Missouri [Terry Tabor, Missouri Department of Health and Senior Services, personal communication, January 28, 2003], certain state health departments expressed interest in accessing the RODS source code. Giving the software away without providing technical support soon proved insufficient. Using the RODS software requires expertise in database, network, geographic information system (GIS), HL7, and system management, capabilities not widely available at that time. Users made multiple requests for customization, support, and assistance with installations, for which resources were not available. Therefore, in September 2003, the University of Pittsburgh released the RODS software under an open-source license, thereby creating the RODS Open Source Project to catalyze the sharing of knowledge and skills related to the software, including its design, installation, configuration, and customization. Materials and MethodsThis section describes the RODS Open Source Project, including the particular license under which RODS is distributed, the infrastructure created to enable growth of the RODS open-source community, methods for publicizing the project and recruiting developers, and the metrics collected to assess its progress. GNU General Public LicenseRODS is distributed as open-source software under the GNU General Public License (GPL) (17), the same open-source license under which Linux® is distributed (18). Unlike the license under which RODS was initially released in December 2002, GPL permits anyone to use, copy, and modify RODS freely. GPL allows consultants and companies to use, install, support, and customize RODS and permits these entities to redistribute their enhanced versions of RODS, provided they make the source code available. This requirement fosters continuous software improvement, benefiting all users and preventing companies from creating proprietary, closed-source versions of RODS. Support for Developers and UsersTo coordinate community-based development of the code, the RODS Laboratory organized the Open Source Project. The RODS modules were classified into six functional areas: data collection, syndrome classification, data warehousing, database encapsulation, outbreak detection, and user interface. Specialists from the laboratory's research and development group named development leaders for each functional area. These development leaders are responsible for recommending new features based on user requests and evaluating whether a developer has the qualifications to contribute source code. Online resources were created to support the Open Source Project, including the RODS Laboratory website (http://www.health.pitt.edu/rods) and a project website hosted on Sourceforge™ (http://openrods.sourceforge.net). The latter site provides standard software project management tools (a concurrent versions system server and patch submission area enabling developers to contribute code), e-mail lists enabling developers and users to communicate, a software-bug reporting system, contact information for the development leaders, and source code for stable versions of the system. Recruitment of Developers and UsersE-mail announcements were sent to 181 persons who had previously downloaded the byte-compiled releases and to all 226 users in the United States who held passwords to the RODS ASP system. Users were given an opportunity for a face-to-face meeting with the core developers at two national conferences, the 2003 National Syndromic Surveillance Conference in New York City and the 2003 American Medical Informatics Fall Symposium in Washington, D.C. Project leaders of other computer-based surveillance projects were also invited. MetricsThe following metrics are collected monthly to manage the project and assess its progress:
The number of installations and the number of contributing developers are considered the two most important metrics. ResultsCurrent Software Architecture of RODS Version 2.0 and Features in DevelopmentA complete technical description of RODS has been published (8). This section describes the system's software architecture and how the modules that comprise that architecture can be used to accomplish different surveillance tasks. RODS 2.0 consists of >42,000 lines of Java code contributed by a team of eight programmers. RODS is a modular system that adheres to CDC's National Electronic Disease Surveillance System (NEDSS) (19) and Public Health Information Network (PHIN) (20) standards so that any of the components can be incorporated into a foreign surveillance system or used to create a native end-to-end RODS system. The RODS software architecture consists of six functional areas: data collection, syndrome classification, data warehousing, database encapsulation, outbreak detection, and user interface (Figure 2). Within the following categories, additional modules are being developed under the Open Source Project (Table 1):
Certain state health departments have requested Lightweight Directory Access Protocol (LDAP) support to enable the creation of seamless links between existing state surveillance systems and the surveillance functions provided by RODS; outside development of such a module is encouraged. State, local, or national health departments can use RODS modules to collect, analyze, and view hospital surveillance data and to view OTC medication sales data from NRDM. A health department can use a subset of these modules to accomplish a specific surveillance task (e.g., receiving and processing free-text chief complaints from hospitals), or it can use all of them (with the RODS database, analytic modules, and user interface) to create an end-to-end surveillance solution. (Examples of how health departments can mix and match RODS modules for different surveillance tasks are available at http://openrods.sourceforge.net.) Project MetricsA total of 480 e-mail announcements about the RODS Open Source Project were sent during the first 3 months of the project. This publicity generated 5,370 page views of the project website, 59 downloads of the source code, and 14 new members to the project mailing lists. One additional installation is using the open-source version of RODS. To date, users are more interested in using the software "as is" and less interested in collaborative feature development. For example, users have asked when the ICD-9 classifier module will be released or whether the system yet works with Microsoft SQL Server.™ Developers at the RODS Laboratory contributed four new features (drilldown of age and sex, customized jurisdictions, a simplified GIS interface, and user preferences) (Table 2). However, at least one health department and one consulting company have expressed interest in collaborating to develop a module that will import XML data into RODS. DiscussionThe goal of the RODS Open Source Project is to accelerate the deployment of computer-based outbreak and disease surveillance systems by writing high-quality surveillance software and catalyzing the formation of a community of users, developers, consultants, and scientists. In the initial years of computer-based outbreak and disease surveillance system development, the main barriers to deployment appeared to be doubts about its efficacy, cost of the technology, concerns about the cost and effect of false alerts on the practice of public health, and legal and administrative issues (25,26). Basic research about data and detectability has been conducted to address concerns about efficacy (2,3,27--29). To address concerns about the effects of false alerts, the RODS laboratory has deployed systems and discovered that persons working in health departments could incorporate the output of these systems into their workflows (4,7). The deployments also established that the cost and effort of deployment is much lower than expected. Finally, the deployments demonstrated that certain concerns about privacy could be addressed. The Health Information Portability and Accountability Act of 1996 (HIPAA), which had not yet become law, nevertheless had a substantial inhibitory effect on hospitals and other covered entities that had data needed by the project. The enactment of the final privacy rule, precedents set by system deployments (4,30--32), and new state laws have helped address certain concerns of data providers (33). Open-source projects can create a community of like-minded persons --- scientists, programmers, consultants, and users --- who have the vision of creating innovative, well-supported software. The importance of catalyzing such a community cannot be overstated. It can strengthen the position of information technology (IT) managers and public health officials who wish to deploy computer-based surveillance systems during planning deliberations. They will be able to assure their supervisors that source code is available, that a pool of developers and consultants exists who can be hired to support the health department if needed, and that ongoing projects in other health departments can help them predict project costs and set appropriate timelines. The RODS Open Source Project enables public health professionals to have a greater role in developing IT solutions to the problem of early detection. Just as public health researchers publish their results in scientific journals, so can they contribute publicly available IT solutions to the RODS Open Source Project. This role might become more apparent as public health personnel become increasingly knowledgeable about public health informatics and work more closely with IT subcontractors and consultants. Continued goals for the RODS Open Source Project are to increase the number of deployments, developers, and supporters of the software. The proposed path for RODS software development is to increase the number of data types the system can accept and implement a range of high-performance outbreak-detection algorithms. One consulting company and one health department have separately expressed interest in collaboratively developing an XML module that can parse non-RODS data sources. The RODS Laboratory and its collaborators at the Auton Laboratory will continue to develop outbreak-detection algorithms (e.g., the wavelet-detection module and WSARE, respectively). ConclusionThe RODS Open Source Project is making software modules available that span the spectrum of processing tasks involved in public health surveillance. Through open source, the project hopes to accelerate the deployment of real-time public health surveillance by lowering costs, increasing reliability, preventing vendor lock-in, and ensuring software customizability. By catalyzing the formation of a community of open-source public health surveillance software advocates, this approach will result in a high-quality software product that achieves mainstream acceptance. Acknowledgments The RODS Open Source Project is supported by the Pennsylvania Department of Health Bioinformatics Grant ME-107. References
Table 1 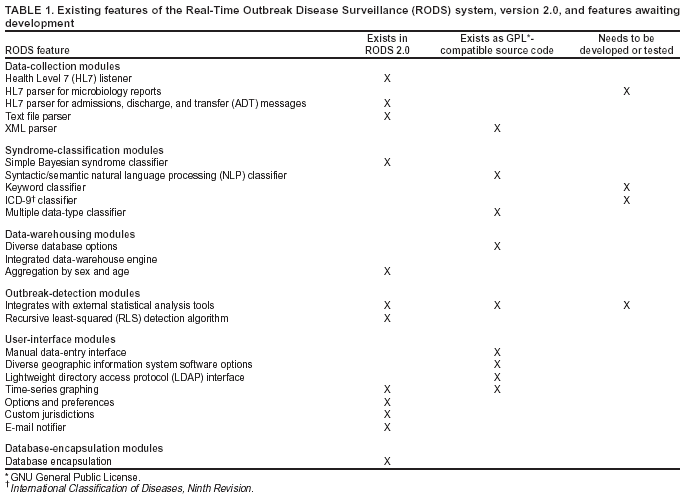 Return to top. Figure 1 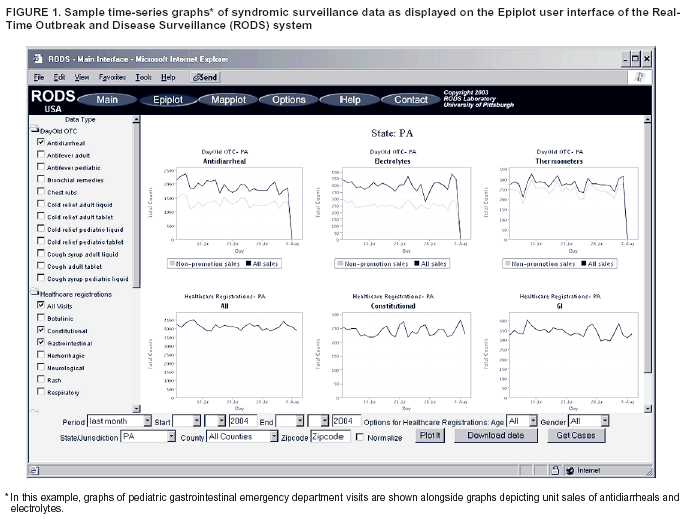 Return to top. Table 2 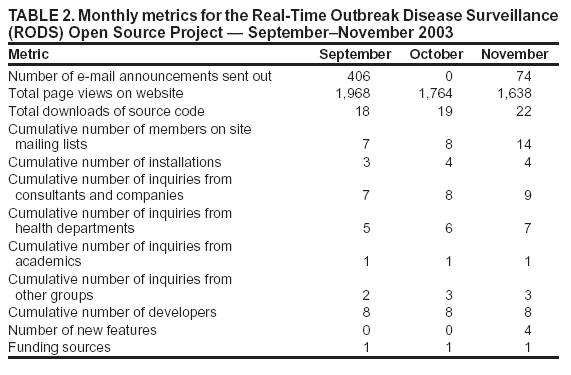 Return to top. Figure 2 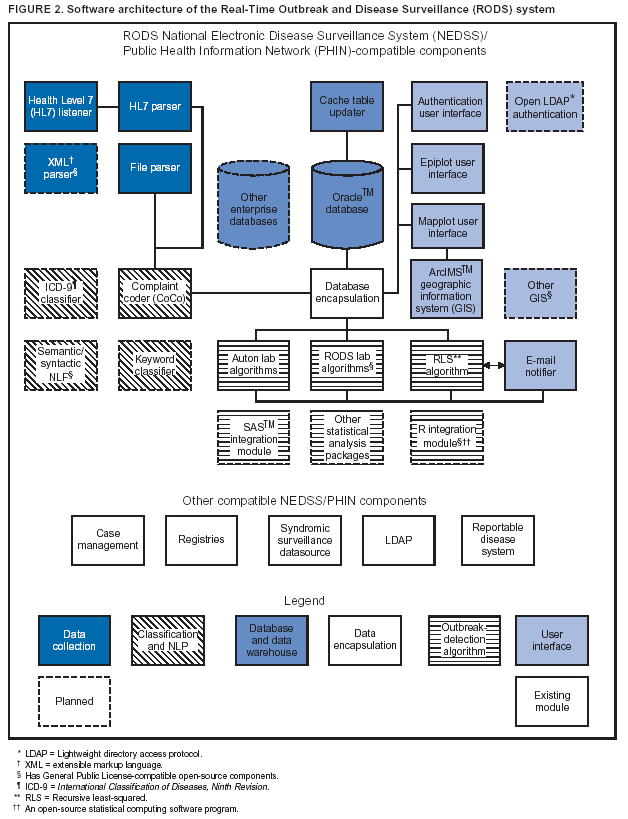 Return to top.
Disclaimer All MMWR HTML versions of articles are electronic conversions from ASCII text into HTML. This conversion may have resulted in character translation or format errors in the HTML version. Users should not rely on this HTML document, but are referred to the electronic PDF version and/or the original MMWR paper copy for the official text, figures, and tables. An original paper copy of this issue can be obtained from the Superintendent of Documents, U.S. Government Printing Office (GPO), Washington, DC 20402-9371; telephone: (202) 512-1800. Contact GPO for current prices. **Questions or messages regarding errors in formatting should be addressed to mmwrq@cdc.gov.Page converted: 9/14/2004 |
|||||||||
This page last reviewed 9/14/2004
|