 |
|
|
|
|
|
|
| ||||||||||
|
|
|
|
|
|
|
||||
| ||||||||||
|
|
|
|
|
Persons using assistive technology might not be able to fully access information in this file. For assistance, please send e-mail to: mmwrq@cdc.gov. Type 508 Accommodation and the title of the report in the subject line of e-mail. Assessment of Susceptibility Testing Practices for Streptococcus pneumoniae --- United States, February 2000Streptococcus pneumoniae is the leading cause of community-acquired pneumonia, otitis media, and meningitis in the United States. Antimicrobial susceptibility results are important for guiding therapy decisions and monitoring emerging resistance patterns. Appropriate methods for pneumococcal susceptibility testing are recommended by the National Committee for Clinical Laboratory Standards (NCCLS) (1--3). Recommendations for pneumococcal susceptibility testing are reviewed annually and were the same in 2000 and 2001. To assess laboratory practices for Streptococcus pneumoniae susceptibility testing on sterile site isolates, in February 2000, CDC conducted a multistate survey of clinical laboratories. This report summarizes the survey results, which found that most practices of clinical laboratories were consistent with NCCLS recommendations; however, some inconsistencies were noted. As antimicrobial resistance in pneumococci continues to worsen, clinical laboratories should be aware of emerging resistance patterns and follow new recommendations to provide clinicians with precise information about antimicrobial susceptibility. Laboratories were selected on the basis of their participation in CDC's Emerging Infections Program/Active Bacterial Core Surveillance (4), through which, since 1995, state and local health departments and universities have conducted active population- and laboratory-based surveillance for invasive pneumococcal disease (defined as isolates from sterile sites such as blood and cerebrospinal fluid [CSF]) in seven to nine geographic areas in the United States. The survey was designed to assess 1) which susceptibility testing practices were being used by clinical laboratories, 2) whether practices followed current NCCLS guidelines, 3) which antimicrobials were being tested routinely, and 4) how microbiology laboratories were reporting susceptibility results to clinicians. A standardized survey was sent to 659 laboratories, and 547 (83%) laboratories responded. A total of 452 (83%) laboratories reported that they tested susceptibility of pneumococcal isolates either in their own laboratory (in-house) or at a reference laboratory, 353 (78%) of which reported doing some in-house testing. Of these 353 laboratories, 188 (53%) performed in-house oxacillin screening on sterile site isolates (Table 1); of these, 187 (99%) followed positive screens with confirmatory minimum inhibitory concentrations (MICs) or had disk diffusion (DD) testing for antimicrobials other than oxacillin. Of the 165 laboratories that bypassed initial oxacillin screening as recommended by NCCLS for blood and CSF isolates, 145 (88%) laboratories performed MICs or DD testing in-house, and the remaining 20 laboratories used a combination of testing in-house and at a reference laboratory. Of the 250 (71%) laboratories that performed MICs or DD testing in-house, 232 (93%) tested sterile site pneumococcal isolates for resistance to penicillin, and 227 (91%) tested a third-generation cephalosporin (cefotaxime or ceftriaxone) (Table 2). In addition, 190 (76%) laboratories tested the three antimicrobials (penicillin, cefotaxime/ceftriaxone, and vancomycin) recommended by NCCLS for blood and CSF isolates, and seven laboratories tested meropenem in addition to these three antimicrobials. Most laboratories also tested sterile site isolates against erythromycin (79%), trimethoprim-sulfamethoxazole (62%), tetracycline (57%), and chloramphenicol (53%); 98 (39%) laboratories tested for resistance to one or more fluoroquinolones. Most laboratories reported using the Etest® (Solna, Sweden) for penicillin (52%) and cefotaxime/ceftriaxone (51%) and disk diffusion for fluoroquinolones (51%); the broth microdilution method was used more frequently (43%--71%) for other antimicrobials. Of the 250 laboratories that performed MICs or DD testing in-house, 207 (83%) laboratories reported susceptibility results to clinicians as interpretations (i.e., susceptible, intermediate, or resistant [S/I/R]), 175 (70%) laboratories reported an exact MIC value, 12 (5%) laboratories reported by zone diameter, and 142 (57%) laboratories used a combination of these reporting methods. A total of 137 (55%) laboratories reported both interpretations and exact MIC values as recommended by NCCLS; however, 66 (26%) laboratories reported only the interpretations, and 35 (14%) laboratories reported only the exact MIC values. Reported by: NL Barrett, MPH, Connecticut Emerging Infections Program; A Reingold, MD, California Emerging Infections Program; K Gershman, MD, Colorado Emerging Infections Program; K McCombs, MPH, Georgia Emerging Infections Program; LH Harrison, MD, Maryland Emerging Infections Program; SK Johnson, MT, Minnesota Emerging Infections Program; JR Hibbs, MD, New York Emerging Infections Program; M Cassidy, P Cieslak, MD, Oregon Emerging Infections Program; A Craig, MD, Tennessee Emerging Infections Program; JH Jorgensen, PhD, Univ of Texas Health Sciences Center, San Antonio. DR Feikin, MD, CG Whitney, MD, Div of Bacterial and Mycotic Diseases and Active Bacterial Core Surveillance/Emerging Infections Program Network, National Center for Infectious Diseases; I Chuang, MD, EIS Officer, CDC. Editorial Note:This survey assessed consistency between reported practices in surveyed laboratories and NCCLS recommendations about oxacillin disk screening, acceptable MIC testing methods and reporting, and antimicrobial agents tested. Most clinical laboratories surveyed were using appropriate methods for pneumococcal susceptibility testing; however, some inconsistencies with NCCLS guidelines were found. In the United States, Streptococcus pneumoniae causes an estimated 63,000 invasive infections and 6,100 deaths per year (4). Since the emergence of penicillin-resistant isolates in the United States in the early 1990s, a high proportion of pneumococci has become resistant to multiple antimicrobial agents. In 1998, approximately 25% of pneumococcal isolates had decreased susceptibility to penicillin, and 14% were resistant to three or more classes of antimicrobial agents (5). The increase in resistance to antimicrobials used to treat pneumococcal infections has resulted in changes in recommended empiric treatment regimens for otitis media, meningitis, and pneumococcal pneumonia (6--8). Initial oxacillin disk screening for pneumococcal isolates is not recommended when isolates come from patients with a potentially life-threatening infection (e.g., meningitis or sepsis). This survey found that 53% of laboratories conducted oxacillin screening on isolates from sterile sites. In the absence of information about the clinical severity of a patient's illness, laboratories should test all isolates from CSF and blood by bypassing oxacillin disk screening and using a more reliable MIC method. Otherwise, definitive MIC results will be delayed by >24 hours, which might prolong use of broad-spectrum antimicrobials chosen for initial empiric treatment. For isolates from other sites (e.g., respiratory), initial oxacillin disk screening is acceptable; however, if the oxacillin zone size is <20 mm, MICs for penicillin and other agents should be determined. Acceptable MIC methods differ for different classes of antimicrobial agents. For ß-lactam agents other than oxacillin, reliable MIC methods include broth microdilution or Etest®. Disk diffusion testing is unreliable for ß-lactam agents including penicillins, cephalosporins, and carbapenems. Either MIC (broth microdilution, Etest®) or disk diffusion should be used for other antimicrobials (e.g., vancomycin, macrolides, trimethoprim-sulfamethoxazole, clindamycin, tetracycline, and fluoroquinolones). If an MIC is determined for an isolate, the exact MIC results should be reported in combination with interpretations (i.e., S/I/R) to assist clinicians with therapeutic decisions, which might vary based on clinical syndrome and severity of illness (3). Antimicrobial choices used for susceptibility testing should include the agents that clinicians use to treat common pneumococcal syndromes. Laboratories should conduct susceptibility testing of all isolates from blood or CSF directly against penicillin, cefotaxime or ceftriaxone, and vancomycin. Meropenem testing also might be performed depending on local clinician preferences and institutional formularies. Because many clinicians use fluoroquinolones as first-line treatment for community-acquired pneumonia or bacteremia, laboratories should perform susceptibility testing against fluoroquinolones. For isolates from patients whose diseases are not life-threatening, such as from middle ear fluid or joint fluid, NCCLS recommends that laboratories perform susceptibility testing for macrolides, trimethoprim-sulfamethoxazole, clindamycin, tetracycline, and fluoroquinolones. Other authorities have recommended that laboratories test against a more extensive primary antimicrobial panel comprising penicillin, cefotaxime or ceftriaxone, and erythromycin, doxycycline or tetracycline, clindamycin, and fluoroquinolones, with trimethoprim-sulfamethoxazole and vancomycin as optional (7). The findings in this report are subject to at least four limitations. First, the survey did not address testing methods used for nonsterile site isolates. Second, the survey assessed laboratory practices in 2000, which might not reflect current practices. Third, the survey assessed reported rather than actual practices. Finally, these laboratories were part of an ongoing surveillance system and might be more likely than other laboratories to be aware of and follow current recommendations. As the problem of antimicrobial resistance for pneumococci worsens, recommendations for susceptibility testing will change, and having precise information on antimicrobial susceptibility will be even more important to clinicians. Clinical laboratories should be aware of new recommendations and emerging resistance patterns. Conducting comprehensive susceptibility testing will enhance the work of public health agencies in tracking emerging resistance patterns in their communities. Acknowledgments This report is based on data contributed by the following members of the Active Bacterial Core Surveillance/Emerging Infections Program Network: L Gelling, MPH, P Daily, MPH, G Rothrock, MPH, D Vugia, MD, California Emerging Infections Program. P Shillam, MSPH, S Burnite, M Finke, MPH, L Hammond, MSPH, Colorado Emerging Infections Program. J Hadler, MD, Connecticut Emerging Infections Program. W Baughman, MS, M Farley, MD, Georgia Emerging Infections Program. MA Pass, J Roche, MD, Maryland Emerging Infections Program. CA Lexau, MPH, R Lynfield, MD, Minnesota Emerging Infections Program. K Stefonek, MPH, Oregon Emerging Infections Program. B Barnes, W Schaffner, MD, Tennessee Emerging Infections Program. L McElmeel, S Crawford, Univ of Texas Health Sciences Center, San Antonio. R Facklam, PhD, T Hilger, C Wright, KA Robinson, MPH, Div of Bacterial and Mycotic Diseases, National Center for Infectious Diseases, CDC. References
Table 1 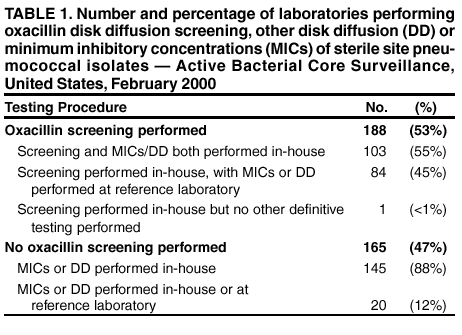 Return to top. Table 2 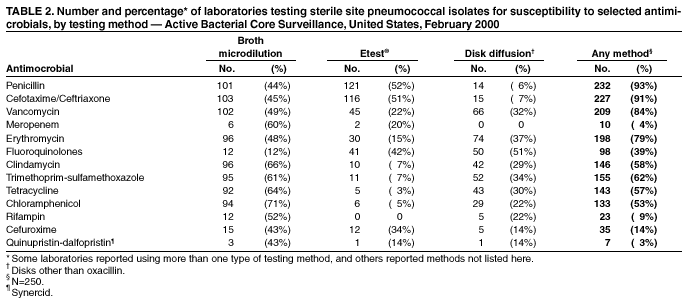 Return to top.
Disclaimer All MMWR HTML versions of articles are electronic conversions from ASCII text into HTML. This conversion may have resulted in character translation or format errors in the HTML version. Users should not rely on this HTML document, but are referred to the electronic PDF version and/or the original MMWR paper copy for the official text, figures, and tables. An original paper copy of this issue can be obtained from the Superintendent of Documents, U.S. Government Printing Office (GPO), Washington, DC 20402-9371; telephone: (202) 512-1800. Contact GPO for current prices. **Questions or messages regarding errors in formatting should be addressed to mmwrq@cdc.gov.Page converted: 5/9/2002 |
|||||||||
This page last reviewed 5/9/2002
|