 |
|
|
|
|
|
|
| ||||||||||
|
|
|
|
|
|
|
||||
| ||||||||||
|
|
|
|
|
Persons using assistive technology might not be able to fully access information in this file. For assistance, please send e-mail to: mmwrq@cdc.gov. Type 508 Accommodation and the title of the report in the subject line of e-mail. Outbreak of Acute Gastroenteritis Attributable to Escherichia coli Serotype O104:H21 -- Helena, Montana, 1994During February-March, 1994, four persons in Helena, Montana (1995 population: 24,569), developed bloody diarrhea and severe abdominal cramps. Stool cultures for Salmonella, Shigella, Campylobacter, and Escherichia coli O157:H7 were negative; however, sorbitol-negative E. coli colonies were identified in stools from all four patients. Isolates from three patients were identified at CDC as a rare serotype -- E. coli O104:H21 that produced Shiga-like toxin II. This report summarizes the epidemiologic and laboratory investigations of this outbreak by the Lewis and Clark County Department of Health and Environmental Sciences, the Montana Department of Health and Environmental Sciences (MDHES), and CDC. A confirmed case was defined as acute infection with E. coli O104:H21 during February 20-May 25, 1994 -- based on stool culture or serologic evidence -- in a resident of or a visitor to the Helena area. A suspected case was defined onset of bloody diarrhea or abdominal cramps during the same period in a resident of or visitor to the Helena area. MDHES and county health departments contacted clinicians, laboratories, and the public through news media reports and requested that suspected cases be reported. Eleven confirmed and seven suspected case-patients were identified Figure_1. Manifestations included abdominal cramps (18 {100%}), diarrhea (17 {94%}), bloody stools (16 {89%}), vomiting (10 {56%}), and fever (six of 15 {40%} for whom information was available). The median age was 36 years (range: 8-63 years), and 12 (67%) were female. Four (22%) persons were hospitalized. Potential sources and risk factors for illness were assessed by a case-control study that included 17 case-patients and three age-, sex-, and neighborhood-matched controls for each case-patient. A history of milk consumption during the 7 days before illness was reported by all 17 case-patients compared with 40 (83%) of 48 * controls (matched odds ratio {OR}=undefined). One brand of milk (Brand A) was significantly associated with illness: of those persons who drank milk at home, 11 (92%) of 12 case-patients compared with 17 (47%) of 36 controls reported drinking Brand A (matched OR=16.0; 95% CI=1.3-492.7). Within this brand, no specific type of milk product was associated with illness. Factors not associated with illness included consumption of other brands of milk, other foods or drinks, and dining in specific restaurants. On May 16, the local and state health departments, the Food and Drug Administration, and CDC inspected the dairy plant where Brand A milk was produced. Based on review of the plant's records for internal microbiologic quality-control testing, on 12 days during February 1-May 13, 1994, the coliform count exceeded the state regulation limiting maximum coliform levels in milk products to less than or equal to 10 coliforms per 100 mL on at least one ready-for-sale milk product. Cultures from selected post-pasteurization piping and equipment surfaces in contact with finished milk products yielded fecal coliforms; however, E. coli O104:H21 was not isolated from any culture samples obtained at the dairy. Two farms provided raw milk for this dairy; rectal swabs obtained from a sample of cattle from these farms did not yield E. coli O104:H21. Reported by: K Moore, Lewis and Clark County Dept of Health and Environmental Sciences; T Damrow, PhD, State Epidemiologist, Montana Dept of Health and Environmental Sciences; DO Abbott, PhD, Montana State Public Health Laboratory. S Jankowski, Microbiology Dept, St. Peter's Community Hospital, Helena. Foodborne and Diarrheal Diseases Br, Div of Bacterial and Mycotic Diseases, National Center for Infectious Diseases, CDC. Editorial NoteEditorial Note: Shiga-like toxin-producing E. coli (SLTEC) are well-recognized causes of gastrointestinal illness, including both bloody and nonbloody diarrhea. E. coli O157:H7, the most common SLTEC, was recognized as a human pathogen in 1982 during the investigation of two outbreaks of bloody diarrhea associated with consumption of commercially sold hamburgers (1). In addition to causing bloody diarrhea, E. coli O157:H7 is the most common cause of hemolytic uremic syndrome (HUS) in children. Although other SLTECs also have been identified in sporadic cases of diarrhea and HUS, the findings in this report document the first reported outbreak of a non-O157 SLTEC in the United States, and the first documentation of illness attributable to Shiga-like toxin-producing E. coli O104:H21. The clinical manifestations of infection in this outbreak were similar to those reported for patients infected with E. coli O157:H7 (2). Although HUS is a well-recognized complication of E. coli O157:H7 infection, no patients developed HUS in this outbreak, possibly reflecting the limited size of the outbreak and the age distribution of patients. Although most outbreaks of E. coli O157:H7 infection have been associated with consumption of ground beef, raw milk also transmits this pathogen (3). Healthy cattle may serve as a reservoir for E. coli O157:H7 and other serotypes of SLTEC (4). The implication of milk in the outbreak in Montana suggests that cows were the original source of this specific strain of E. coli O104:H21. Although the investigation documented post-pasteurization contamination of milk products with fecal coliforms, E. coli O104:H21 was not isolated from cultures obtained at the dairy, possibly because not all post-pasteurization equipment surfaces were sampled or because of the absence of the pathogen within the dairy at the time of the inspection. Because the techniques used to identify non-O157 SLTEC are not available in most laboratories (3), infections caused by this pathogen are most likely to be unrecognized. Most clinical laboratories that test for E. coli O157:H7 screen stools on a special medium (sorbitol-MacConkey agar {SMAC}) because E. coli O157:H7 isolates do not ferment sorbitol after overnight incubation (5), and most laboratories routinely discard sorbitol-positive colonies and sorbitol-negative colonies that do not agglutinate in O157 antiserum. Therefore, isolates of E. coli O104:H21 and other non-O157 SLTEC are not recognized. The increased availability in clinical laboratories of techniques such as testing for Shiga-like toxin or the genes encoding this protein may enhance the detection of disease attributable to non-O157 SLTEC. When evaluating clusters of patients with bloody diarrhea and other severe diarrheal illness, health-care providers also should consider the potential roles of E. coli O104:H21 or another non-O157 SLTEC. When cultures of stool are negative for specific pathogens, the state health department can be contacted to determine whether specimens should be examined further for SLTEC. When advised, health-care providers should freeze fecal specimens and store isolates from patients with bloody diarrhea; such specimens may assist in a subsequent investigation. References
* Persons who responded "Don't know" to any question were excluded from the analysis. Figure_1 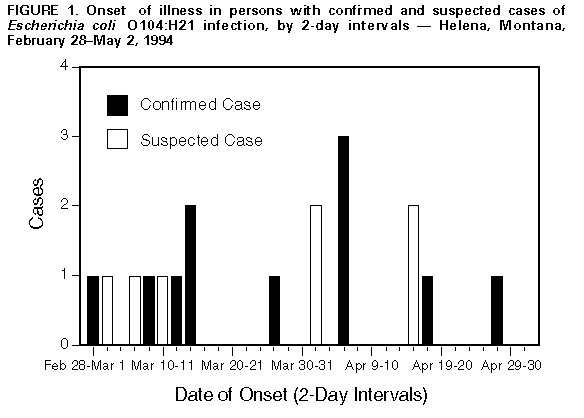 Return to top. Disclaimer All MMWR HTML versions of articles are electronic conversions from ASCII text into HTML. This conversion may have resulted in character translation or format errors in the HTML version. Users should not rely on this HTML document, but are referred to the electronic PDF version and/or the original MMWR paper copy for the official text, figures, and tables. An original paper copy of this issue can be obtained from the Superintendent of Documents, U.S. Government Printing Office (GPO), Washington, DC 20402-9371; telephone: (202) 512-1800. Contact GPO for current prices. **Questions or messages regarding errors in formatting should be addressed to mmwrq@cdc.gov.Page converted: 09/19/98 |
|||||||||
This page last reviewed 5/2/01
|