COVID-19 Science Update released: April 24, 2020 Edition 7

The COVID-19 Science Update summarizes new and emerging scientific data for public health professionals to meet the challenges of this fast-moving pandemic. Weekly, staff from the CDC COVID-19 Response and the CDC Library systematically review literature in the WHO COVID-19 databaseexternal icon, and select publications and preprints for public health priority topics in the CDC Science Agenda for COVID-19 and CDC COVID-19 Response Health Equity Strategy.
Here you can find all previous COVID-19 Science Updates.
PEER-REVIEWED
Characteristics of pediatric SARS-CoV-2 infection and potential evidence for persistent fecal viral sheddingexternal icon. Xu et al. Nature Medicine (March 13, 2020).
Key findings:
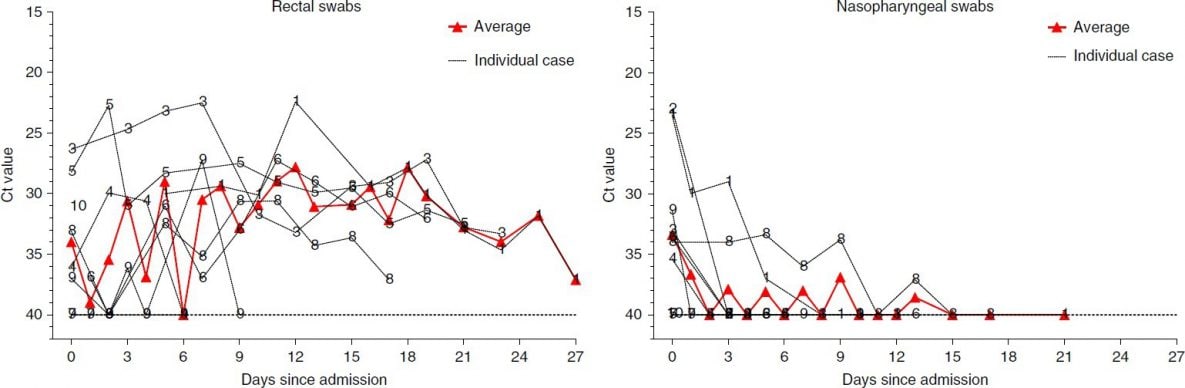
- Among 10 pediatric COVID-19 patients who initially tested negative for SARS-CoV-2 using nasopharyngeal swabs, 8 subsequently tested positive using rectal swabs (Figure).
- RT-PCR results suggested greater viral shedding in the digestive tract than the respiratory tract, especially after the first week post-admission.
Methods: A prospective observational study of 10 pediatric COVID-19 patients (aged 2 months – 15 years) hospitalized in Guangzhou, China. Serial RT-PCR performed to detect viral excretion from the respiratory and gastrointestinal tracts. Limitations: No assessment for replication-competent virus; small single-center study.
Implications: Children appear to shed SARS-CoV-2 RNA in their stool longer that in the nasopharynx.
Figure:
Note: From Xu Y et al. The Ct values shown on both y-axes to quantify viral load in the rectal and nasopharyngeal samples of the 10 pediatric patients. Lower Ct values indicate more virus. The red line shows the average Ct value. Available via Nature Public Health Emergency Collection through PubMed Central.
Sequential analysis of viral load in a neonate and her mother infected with SARS-CoV-2external icon. Han et al. Clinical Infectious Diseases (April 16, 2020).
Key findings:
- All samples from a neonate (respiratory tract, stool, urine, and blood) tested positive for SARS-CoV-2.
- Only respiratory and stool samples were positive in the mother, and viral load was low (mother’s breast milk tested negative).
- The neonate had a fever and gastrointestinal symptoms, while the mother did not.
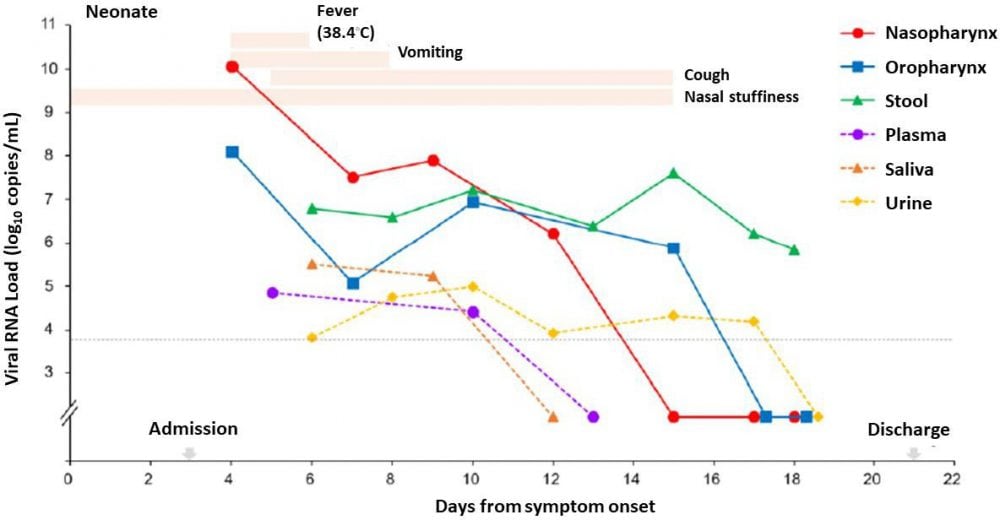
- High levels of virus persisted in the neonate’s stool after all other samples tested negative and the neonate’s gastrointestinal symptoms improved (Figure).
Methods: Case-report of a mother and her 1-month old female neonate hospitalized in Seoul, South Korea. Samples from multiple sites were tested by RT-PCR to quantify SARS-CoV-2 viral load, and patients’ clinical manifestations of COVID-19 were assessed for 3 weeks. Limitations: No assessment for replication-competent virus; mode of neonatal infection uncertain but there was close contact with the infected mother during breast-feeding (breast milk tested negative for virus) and infection occurred within a family cluster.
Implications: A neonate shed SARS-CoV-2 RNA in stool longer than detectable RNA was shed in other anatomic sites including the respiratory tract.
Figure:
Note: Adapted from Han et al. The neonate’s symptoms, and their duration, are shown as peach bars at the top of the figure. The green line shows viral RNA load (log10 copies/mL on the y-axis) over time in the stool, which remained high after all other samples from the nasopharynx, oropharynx, plasma, saliva, and urine tested negative. Adapted and reproduced by permission of Oxford University Press on behalf of the Infectious Diseases Society of America. Please visit: https://academic.oup.com/cid/advance-article/doi/10.1093/cid/ciaa447/5820869external icon. OUP and the IDSA are not responsible or in any way liable for the accuracy of the adaptation. Licensee is solely responsible for the adaptation in this publication/reprint.
PEER-REVIEWED
Thrombocytopenia and its association with mortality in patients with COVID-19external icon. Yang et al. Journal of Thrombosis and Hemostasis (April 17, 2020).
Key Findings:
- 37% of patients had abnormal platelet counts: 21% very low platelet and 16% low.
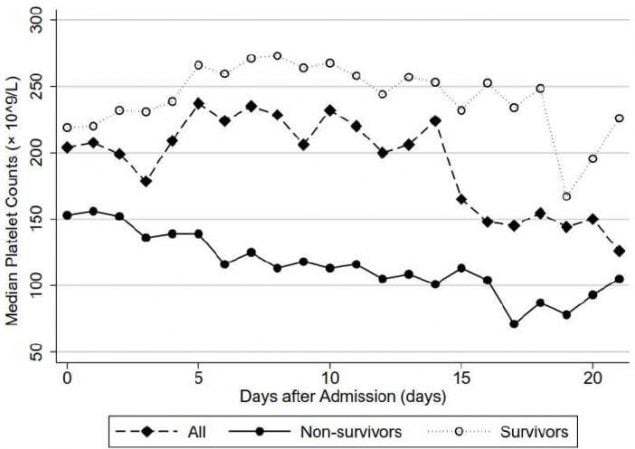
- Patients who died had lower median platelet counts than those who survived (Figure 1).
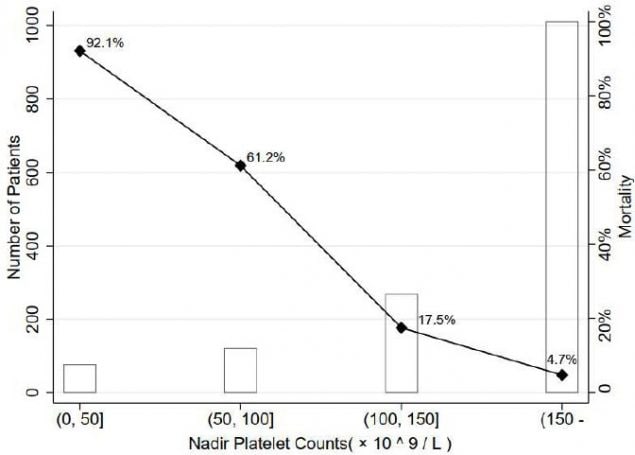
- In-hospital mortality increased as platelet count decreased (Figure 2).
Methods: Observational study of 1,476 hospitalized adults who were either discharged or deceased by February 25, 2020. Investigators retrospectively grouped patients into four categories based on their lowest recorded platelet count and examined the association between platelet count and in-hospital mortality. Limitations: Single-center study; number of platelet count tests varied across patients; potential contribution of other comorbidities or laboratory abnormalities not accounted for in analysis.
Implications: Low platelet counts were associated with higher in-hospital mortality among COVID-19 patients. A causal relationship remains unclear and low platelets may be a marker for severe disease from other causes.
Figure 1
Figure 2
Note: From Yang et al. Figure 1. The three lines in the figure represent: COVID-19 patient survivors (hollow circles, faint line); all participants (diamonds, dotted line), and; non-survivors (black circles, solid line). The y-axis represents median platelet counts (not nadir platelet counts). Non-survivors had lower median platelet counts than survivors during the trajectory of their hospital stay. Figure 2. The black line represents the in-hospital mortality, which is highest (92.1%) in the group with the lowest (i.e., nadir) platelet count (shown on the x-axis). The hollow bars represent the number of patients in each of the four platelet count groups. Used by permission of John Wiley & Sons, Inc. ©International Society on Thrombosis and Haemostasis.
Neurological features in severe SARS-CoV-2 infectionexternal icon. Helms et al. NEJM (April 15, 2020).
Key Findings
- Among 58 COVID-19 patients admitted to ICU with ARDS:
- 14% presented with neurological symptoms.
- 69% developed signs of agitation after sedation and neuromuscular blockers to treat ARDS were discontinued; the majority (65%) showed confusion.
- 67% had diffuse reflex anomalies.
- At discharge, 33% showed inattention, disorientation or poorly organized movements in response to commands.
- Brain magnetic resonance imaging (MRI) for 13 patients with unexplained neurological symptoms found a range of imaging abnormalities, including 3 cases of ischemic stroke.
- Cerebrospinal fluid (CSF) tested negative for SARS-CoV-2 in all 7 patients.
Methods: Observational study of 58 COVID-19 patients hospitalized with ARDS in Strasbourg, France. Neurological assessments were performed on ICU admission and at discharge; additional brain MRIs and CSF analysis were done for selected patients. Limitations: Six patients excluded due to severe illness or death.
Implications: In a small observational study, two-thirds of patients hospitalized with severe COVID-19 illness and ARDS experienced neurological impairment. It is unclear whether some of this impairment is reversible and whether it resulted from infection, host immune response, treatment withdrawal, or some combination.
No SARS-CoV-2 detected in amniotic fluid in mid-pregnancy.external icon Yu et al. Lancet Infectious Diseases (April 22, 2020)
Key findings:
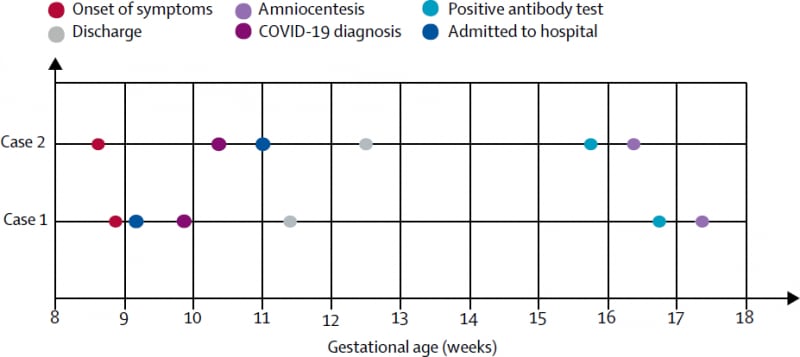
- In two women recovered from COVID-19 during the first trimester of pregnancy who developed positive SARS-CoV-2 serologies, amniotic fluid from amniocenteses during the second trimester was negative for SARS-CoV-2 by RT-PCR and negative for anti-SARS-CoV-2 IgM and IgG.
Methods: Retrospective review of clinical records and laboratory results. Amniotic fluid samples and sera tested for IgM and IgG by serologic tests. Amniotic fluid and throat swabs also tested by RT-PCR. Limitations: Small sample size; lack of cord blood; SARS-CoV-2 may be transient in amniotic fluid; birth outcomes not presently available.
Implications: Two case reports showed no evidence of intrauterine SARS-CoV-2 transmission (ultimate birth outcomes are pending currently).
Figure:
Note: Adapted from Yu et al. Timeline of exposure to SARS-CoV-2 and amniocentesis. This article was published in Lancet Infectious Diseases, Yu et al., No SARS-CoV-2 detected in amniotic fluid in mid-pregnancy, Copyright Elsevier 2020. This article is currently available at the Elsevier COVID-19 resource center: https://www.elsevier.com/connect/coronavirus-information-centerexternal icon.
PREPRINTS (NOT PEER-REVIEWED)
Outcomes of hydroxychloroquine usage in United States veterans hospitalized with COVID-19external icon. Magagnoli et al. medRxiv (April 21, 2020). Published external iconin Med (June 5, 2020).
Key findings:
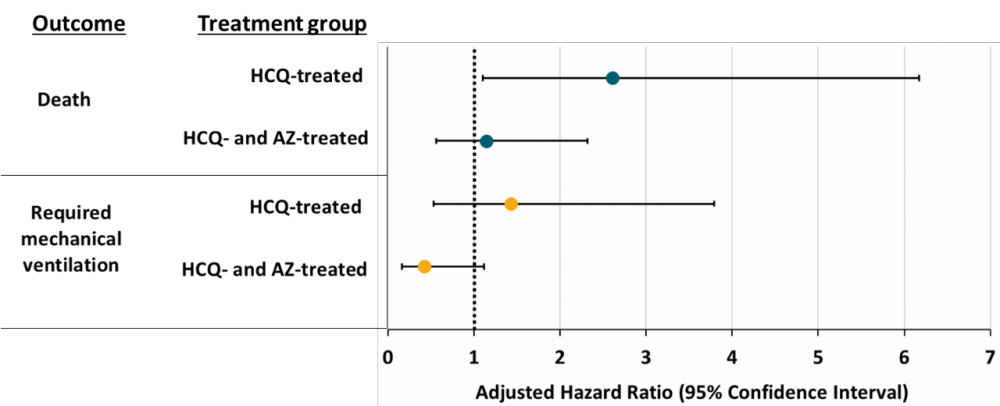
- COVID-19 patients treated with hydroxychloroquine (HCQ) had an increased risk of death compared with patients treated with standard care after adjusting for disease severity (Figure).
- Patients treated with HCQ did not have an increased risk of requiring mechanical ventilation (Figure).
Methods: Nationwide, retrospective cohort study of 393 male COVID-19 patients hospitalized in Veterans Health Administration medical centers between March 9, 2020 and April 11, 2020. Patients all received standard care and were grouped according to additional treatment: (1) HCQ alone (n = 97); (2) HCQ and azithromycin (HCQ-AZ) (n = 113); or (3) no additional treatment (n = 158). Hazard ratios were used to compare the risk of death and mechanical ventilation for patients treated with HCQ alone or with HCQ-AZ compared with no HCQ. Propensity score models were used to account for non-randomized assignment to the treatment groups, and hazard ratios were calculated that adjusted for baseline covariates, including demographics, comorbidities, vital signs, and laboratory values. Limitations: Convenience sample; no safety data reported, entirely male sample.
Implications: Findings from a non-randomized convenience sample suggest that HCQ offered no benefit and may have increased risk of death among COVID-19 patients.
Figure:
Note: Adapted from Magagnoli et al. Adjusted hazard ratios and 95% confidence intervals for patient death and mechanical ventilation in patients in each treatment group compared with those receiving only standard care and no additional treatment. Hazard ratios with 95% confidence intervals not crossing the dotted line (adjusted hazard ratio = 1) are considered statistically significant. HCQ: hydroxychloroquine, AZ: azithromycin. Licensed under CC-BY-NC-ND 4.0.
PEER-REVIEWED
Severe acute respiratory syndrome coronavirus 2−specific antibody responses in coronavirus disease 2019 patients. Okba et al. Emerging Infectious Diseases (April 8, 2020).
Key findings:
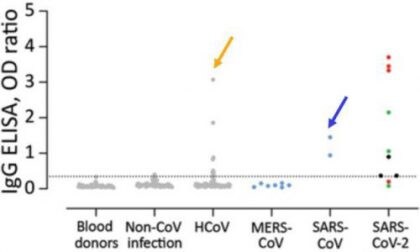
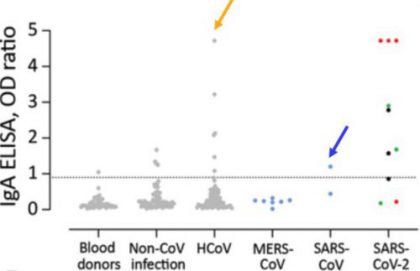
- Commercial tests for anti-SARS-CoV-2 IgG and IgA produced false-positive results when used to test serum samples from patients with other coronavirus infections, including common seasonal human coronaviruses (HCoV) that generally cause of mild human illness (gold arrows) and SARS-CoV-1 (cause of SARS, blue arrows) (Figure).
Methods: Investigators validated commercial tests for SARS-CoV-2 IgG and IgA antibodies (EUROIMMUN Medizinische Labordiagnostika AG) using serum samples from 45 healthy blood donors, 76 patients with non-CoV respiratory infections, 75 with HCoV infections, 7 with MERS-CoV infections, 3 with SARS-CoV-1 infection and 12 with PCR-confirmed SARS-CoV-2 infection. Limitation: Only two commercial assays evaluated.
Implications: Because most people have antibodies against seasonal HCoVs, it is important to ensure SARS-CoV-2 antibody tests will not produce false-positive results because they detect these other (cross-reacting) antibodies.
Figure:
Note: Adapted from Okba et al. Validation of the specificity of commercial ELISAs for detection of IgG (left panel) and IgA (right panel) against SARS-CoV-2 spike protein subunit 1. Gray dots indicate healthy blood donors, non-CoV respiratory infections, and HCoV infections; blue dots indicate non-SARS-CoV-2 zoonotic coronavirus infections (MERS-CoV and SARS-CoV-1); cohort of SARS-CoV-2 infection indicated by red dots (severe infection) and black and green dots (mild infection). Dotted horizontal lines indicate ELISA positive cutoff values. Dots for SARS-CoV-2 below this line were from ≤ 8 days after onset of illness. ELISA=enzyme- linked immunosorbent assay; OD=optical density using a 450 nm filter. Open access journal; all content freely available.
Saliva is a reliable tool to detect SARS-CoV-2external icon. Azzi et al. Journal of Infection (April 14, 2020)
Key Findings
- In a series of 25 patients, saliva samples at hospitalization all had detectable SARS-CoV-2 RNA.
- Of 8 patients from whom a second saliva sample was assayed 4 days later, all remained positive including 2 patients that >2 negative respiratory tract specimens.
Methods: Saliva samples from 25 hospitalized patients with severe COVID-19 were tested for SARS-CoV-2 RNA by RT-PCR. Limitations: Small study limited to patients with severe illness; baseline data with minimal follow-up.
Implications: Salivary samples are a less-invasive specimen that may be at least as sensitive as respiratory samples to detect SARS-CoV-2 RNA early in illness. However, data are not yet conclusive to recommend use of saliva samples as a reliable tool to diagnose and monitor SARS-CoV-2 infection.
SARS-CoV-2 isolation from ocular secretions of a patient with SARS-COV2 in Italy with prolonged viral RNA detectionexternal icon. Colavita et al. Annals of Internal Medicine (April 17, 2020).
Key findings:
- A patient with mild illness but severe conjunctivitis had viraldetectable in conjunctival swabs.
- Viral RNA was detected in conjunctival swabs for up to 27 days.
- Conjunctivitis had resolved by day 21.
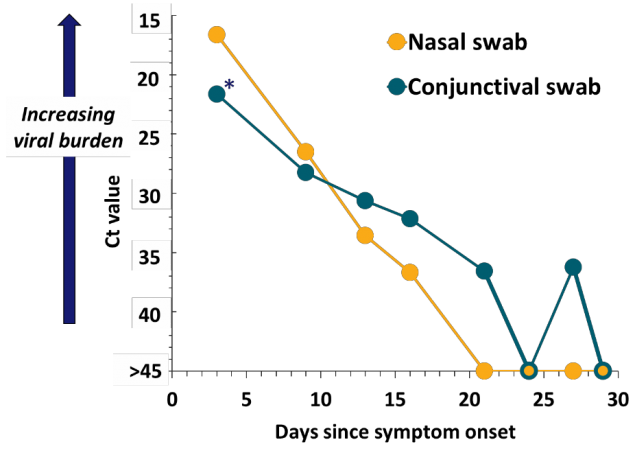
- Viral RNA became undetectable in the nasal swabs after day 16 (Figure).
- Viral culture from day 3 conjunctival swab may have yielded replication-competent SARS-CoV-2.
Methods: Case report of a 65-year-old hospitalized woman. Due to severe conjunctivitis at admission she underwent eye exams and with nasal and conjunctival swabs collections for 29 days. Swabs were tested for RNA by RT-PCR. Limitations: Culture methods did not include serial passage.
Implications: This may be the first report suggesting infectious virus is present in conjunctival secretions.
Figure:
Note: Adapted from Colavita, et al. Figure shows Ct values for SARS-CoV-2 RNA in ocular and nasal swabs by collection data. Asterisk (*) indicates swab from which culturable virus was potentially isolated. Available via American College of Physicians Public Health Emergency Collection through PubMed Central.
- Yancy . COVID-19 and African Americansexternal icon. JAMA. Perspective on how health care disparities, including ability to adequately “social distance,” in the United States are translating into higher rates of infection and case fatality rates among African Americans.
- Artenstein et al. In pursuit of PPEexternal icon. NEJM. Department of Homeland Security and FBI involved in attempted seizure of PPE shipment purchased by Massachusetts hospital.
- Webster P. Virtual healthcare in the era of COVID-19external icon. Lancet. COVID-19 pandemic requires more telemedicine and some clinicians are embracing it.
- El-Sadr et al. Africa in the path of COVID-19external icon. NEJM. Although African countries reported a combined ~14,000 cases as of April 13, challenges are numerous (e.g., ICU beds, ability to social distance) and will require global coordination.
- Bauchner et al. Health Care heroes of the COVID-19 pandemicexternal icon. JAMA. Journal editors salute millions of healthcare workers and recognize astute physician-leaders during COVID-19 pandemic worldwide.
- Metlay et al. Clinical decision making: Weighing evidence to inform clinical decisionsexternal icon. Annals of Internal Medicine. Discusses how to make informed COVID-19 patient care decisions when evidence-base is scant, conflicting, and growing.
- Kaiser J. To streamline coronavirus vaccine and drug efforts, NIH and firms join forcesexternal icon. Science. Describes the ACTIV initiative, a public-private partnership between NIH, pharma, and biotech companies to coordinate efforts to conduct clinical trials of drugs and vaccines.
Disclaimer: The purpose of the CDC COVID-19 Science Update is to share public health articles with public health agencies and departments for informational and educational purposes. Materials listed in this Science Update are selected to provide awareness of relevant public health literature. A material’s inclusion and the material itself provided here in full or in part, does not necessarily represent the views of the U.S. Department of Health and Human Services or the CDC, nor does it necessarily imply endorsement of methods or findings. While much of the COVID-19 literature is open access or otherwise freely available, it is the responsibility of the third-party user to determine whether any intellectual property rights govern the use of materials in this Science Update prior to use or distribution. Findings are based on research available at the time of this publication and may be subject to change.