COVID-19 Science Update released: April 16, 2021 Edition 85

The COVID-19 Science Update summarizes new and emerging scientific data for public health professionals to meet the challenges of this fast-moving pandemic. Weekly, staff from the CDC COVID-19 Response and the CDC Library systematically review literature in the WHO COVID-19 databaseexternal icon, and select publications and preprints for public health priority topics in the CDC Science Agenda for COVID-19 and CDC COVID-19 Response Health Equity Strategy.
Section headings in the COVID-19 Science Update align with the CDC Science Agenda for COVID-19.
Section headings in the COVID-19 Science Update have been changed to align
with the CDC Science Agenda for COVID-19.
Here you can find all previous COVID-19 Science Updates.
PEER-REVIEWED
A pair of case series report on European patients with thrombotic events or intracranial hemorrhage beginning several days after vaccination with ChAdOx1. These studies describe observed patterns.
A. Thrombotic thrombocytopenia after ChAdOx1 nCov-19 vaccination.external icon Greinacher et al. NEJM (April 9, 2021).
Key findings:
- 11 patients presented with one or more thrombotic events beginning 5-16 days after vaccination:
- 9 had cerebral venous thrombosis, 1 had intracranial hemorrhage, and 5 had disseminated intravascular coagulation.
- 6 patients died.
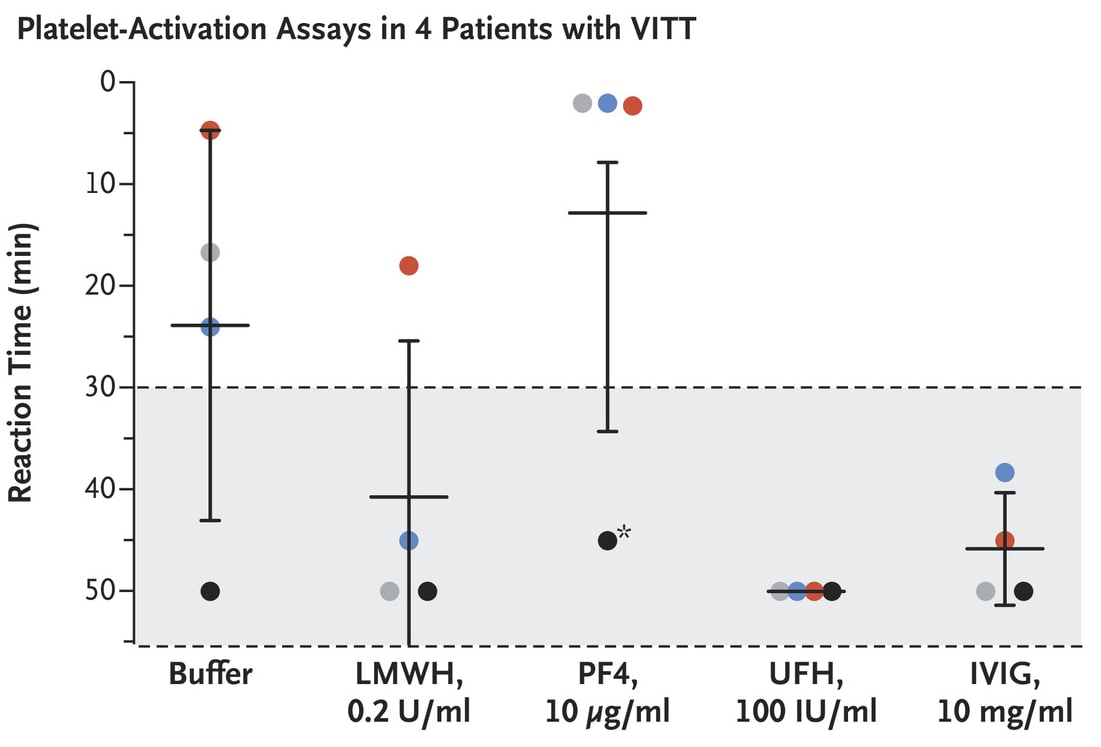
- Serum samples from a subset (n = 4), showed platelet reactivity was inhibited by low-molecular-weight heparin and greatly enhanced by platelet factor 4 (PF4) (Figure).
- None received any heparin prior to presentation.
Methods: Case series of clinical and laboratory features of 11 patients (9 women, median age 36) in Germany and Austria with thrombosis or thrombocytopenia 5-16 days after vaccination with ChAdOx1 nCov-19. Standard ELISA to detect PF4–heparin antibodies and a modified (PF4-enhanced) platelet-activation test were utilized. Limitations: Small sample size.
Figure:
Note: Adapted from Greinacher et al. Platelet-activation assay results in serum samples (each shown by a color) from a subset (n = 4) of patients. Horizontal bars show means and vertical bars are standard errors. Shorter reaction time indicates stronger platelet-activation; reaction time >30 minutes indicates clinically insignificant reactivity. Asterisk indicates an outlier sample, which was strongly positive on additional testing. UFH = unfractionated heparin; IVIG = intravenous immune globulin. From the New England Journal of Medicine, Greinacher et al., Thrombotic thrombocytopenia after ChAdOx1 nCov-19 vaccination. April 9, 2021, online ahead of print. Copyright © 2021 Massachusetts Medical Society. Reprinted with permission from Massachusetts Medical Society.
B .Thrombosis and thrombocytopenia after ChAdOx1 nCoV-19 vaccination.external icon Schultz et al. NEJM (April 9, 2021).
Key findings:
- 5 health care workers (ages 32-54) presented with thrombosis (4 with cerebral venous thrombosis) and thrombocytopenia 7-10 days after 1st dose.
- 3 died.
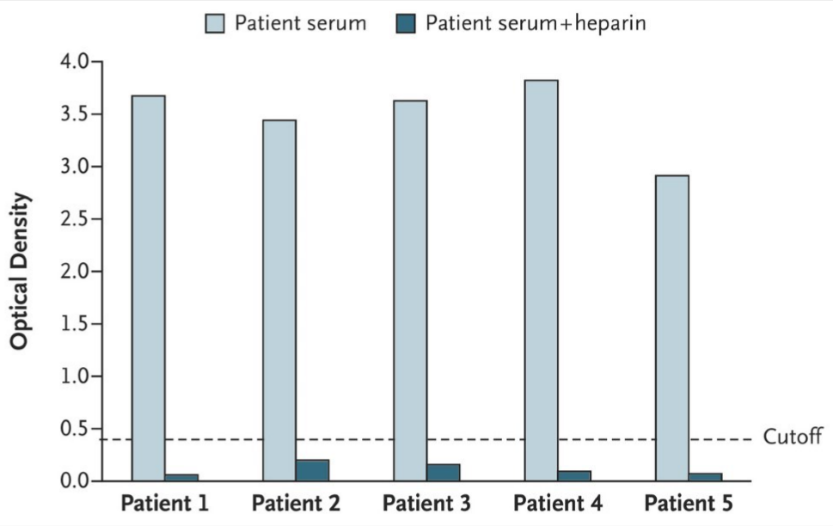
- Despite no previous heparin exposure, all had sera with high antibody levels to platelet factor 4 (PF4)–polyanion complexes that were inhibited by heparin. (Figure)
Methods: In Norway, where ~130,000 people received ChAdOx1, 5 health care workers presented with thrombotic symptoms 7-10 days after receiving a 1st dose. Standard ELISA was used to test for antibodies to PF4 in complex with poly(vinyl sulfonate) (heparin analogue) in patient serum. Confirmatory inhibition tests conducted under high heparin concentration. Limitations: Small sample size.
Figure:
Note: Adapted from Schultz et al. IgG PF4–polyanion antibodies in the patients’ serum samples, measured by enzyme-linked immunosorbent assay, before and after addition of a saturating dose of heparin (100 IU/mL). Mean optical density ≥ 0.4 indicates the presence of antibodies. From the New England Journal of Medicine, Schultz et al., Thrombosis and thrombocytopenia after ChAdOx1 nCoV-19 vaccination. April 9, 2021, online ahead of print. Copyright © 2021 Massachusetts Medical Society. Reprinted with permission from Massachusetts Medical Society.
Implications for both studies (Greinacher et al and Schultz et al): Despite no previous heparin exposure, evidence suggests that in rare cases, ChAdOx1 nCov-19 vaccination can result in immune thrombotic thrombocytopenia mediated by platelet-activating antibodies against PF4.
Comparison of estimated effectiveness of case-based and population-based interventions on COVID-19 containment in Taiwanexternal icon. Ng et al. JAMA Internal Medicine (April 6, 2021).
Key findings:
- Case-based interventions alone decrease the reproduction number (R) to 1.53 (95% credible interval [CrI], 1.50 − 1.57), not sufficient for epidemic control.
- Population-based interventions reduce R to 1.30 (95% Crl, 1.03 − 1.58), also not sufficient for epidemic control.
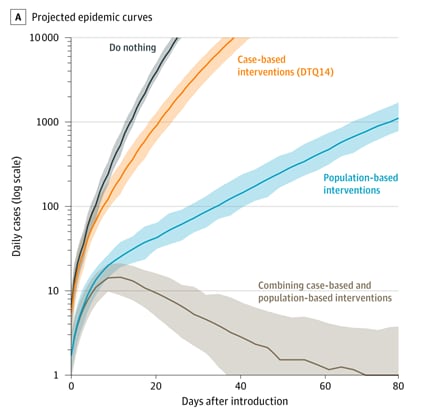
- Both case-based and population-based interventions were needed to reduce the SARS-CoV-2 reproductive rate to <1 in Taiwan (Figure).
Methods: A comparative effectiveness study of case-based (detection, contact tracing, and 14-day quarantine of close contacts) and population-based interventions (use of face masks, personal hygiene, and physical distancing) using a stochastic branching process model of 158 confirmed COVID-19 cases in Taiwan between March 2020 and February 2021. Limitations: Accuracy of the model depends on input parameters; parameters based on Taiwan may not be generalizable.
Implications: Taiwan reported no locally acquired cases of COVID-19 for 253 days between April and December 2020. The combination of case-based and population-based intervention strategies could explain how Taiwan was able to mitigate COVID-19 spread in the local population.
Figure:
Note: Adapted from Ng et al. Projected epidemic curves with 100 initial introductions under 4 different intervention scenarios: (1) Do nothing (no intervention, R = 2.50), (2) case-based interventions (case surveillance, contact tracing, and quarantine) only, (3) population-based interventions (physical distancing and masking) only, and (4) combining case-based and population-based interventions. Licensed under CC BY.
Association of sociodemographic factors and blood group type with risk of COVID-19 in a US population.external icon Anderson et al. JAMA Network Open (April 5, 2021).
Key findings:
- Blood type was not associated with disease susceptibility (viral positivity) or severity (hospitalization or ICU admission):
- Compared with type O blood, type A was not associated with increased viral positivity (aOR 0.97 [95% CI 0.93-1.01]), hospitalization (aOR 0.89 [95% CI 0.80-0.99]), or ICU admission (aOR 0.84 [95% CI 0.69-1.02]).
- Similarly, types B and AB were not associated with worse outcomes compared with type O.
- Analyses restricted to White race produced similar results.
Methods: Case-control study in a large, nonprofit, integrated health care system’s electronic health record database (n = 107,796) was conducted between March 3 and November 2, 2020 to test whether blood type is associated with SARS-CoV-2 susceptibility and COVID-19 severity. Limitations: Study results may not be generalizable to all populations.
Implications: ABO groups were not associated with SARS-CoV-2 susceptibility nor COVID-19 illness severity.
PEER-REVIEWED
Implications of the school-household network structure on SARS-CoV-2 transmission under school reopening strategies in Englandexternal icon. Munday et al. Nature Communications (March 29, 2021).
Key findings:
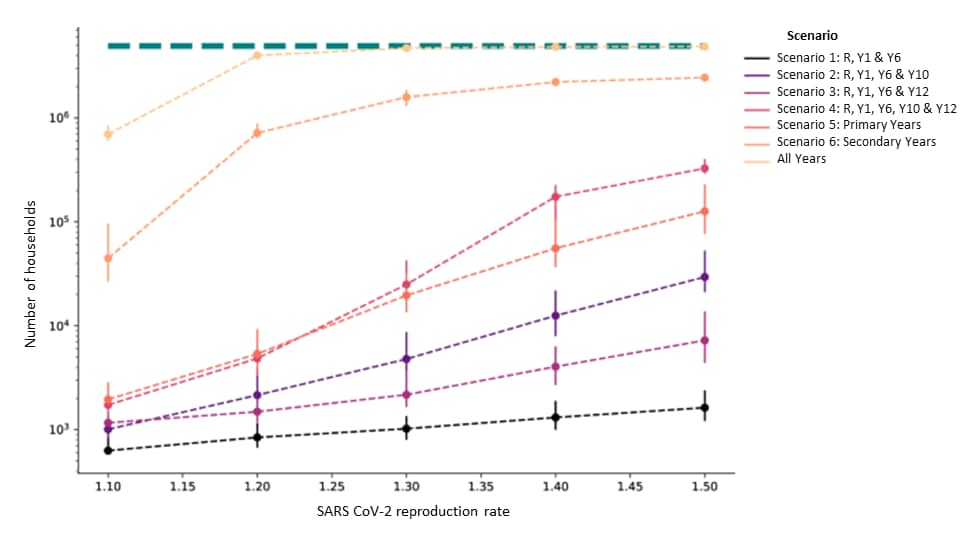
- Scenario 1 (Reception and Years 1 and 6) had the lowest estimated probability (1-10%) for a school infecting another school and put the fewest number of households at risk (Figure).
- Scenario 6 (opening secondary schools only) had the highest estimated probability (25%) of infecting multiple schools and affected the most households.
- When the network only included pupils from certain years, the expected number of schools infected by others in the network decreased.
Methods: Household addresses of school children were used to identify a network of 21,583 schools attended by 4.6 million primary school and 3.4 million secondary school children. Scenarios based on opening different age-based school levels were created (e.g., Reception [ages 4–5], Year 1 [ages 5-6] through Year 12 [ages 16-17]). Probability of transmission between school pairs was estimated in a transmission probability network for each reopening scenario. Limitations: Model does not account for additional mitigation strategies; assumes a well-mixed (homogenous) contact network within each school and no presence of immunity.
Implications: Risk posed by reopening secondary schools differs from the risk posed by reopening primary schools due to the connectivity and potential transmission networks between schools. Reopening strategies should consider cross-school transmission potential.
Figure:
Note: Adapted from Munday et al. Estimated number of households with children in a binary transmission network under different school reopening scenarios and SARS CoV-2 reproduction rates. The Green dashed line shows the total number of households (4,927,163) in the school system. Secondary schools (mean age ≥11 years), Primary schools (mean age ≤11 years), Reception (R, ages 4-5). Points show median and vertical bars are 90% credible intervals. Licensed under CC BY 4.0.
Epidemiologic evidence for airborne transmission of SARS-CoV-2 during church singing, Australia, 2020. Katelaris et al. Emerging Infectious Diseases (April 5, 2021).
Key findings:
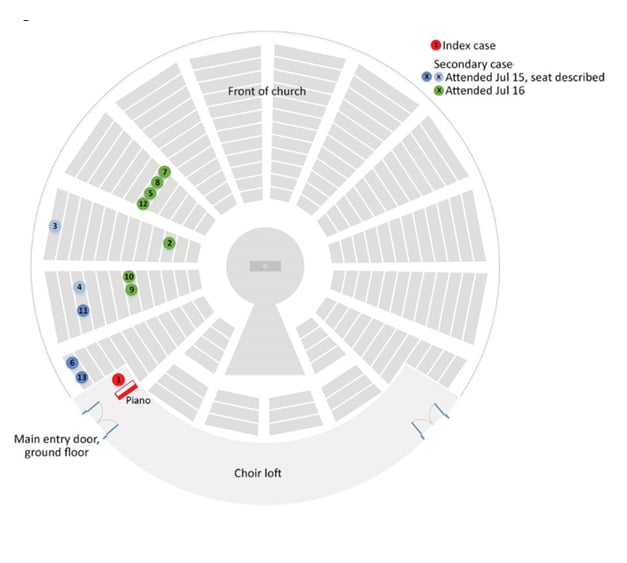
- 18-year old chorist was the index case for an outbreak at a church.
- Chorist sung from the choir loft at four 1-hour services, 3.5 meters above the congregation.
- 12 secondary cases from 2 separate church services, seated up to 15 meters away, were linked to the index case (Figure).
- The index case sung in a space with minimal ventilation. Respiratory particles accumulated and convection currents carried particles to the congregation.
- Masks were not used.
Methods: 508 contacts who attended at least one church service were monitored for 17 days following their exposure date. Seating positions of the congregation were determined via interviews and video footage to confirm location at time of exposure. SARS-CoV-2 genome sequencing was performed to establish the genomic cluster. Limitations: Most contacts were tested within a week of exposure, which could have been too early to detect some asymptomatic infections.
Implications: Additional mitigation measures might be necessary to prevent airborne infection during indoor singing, including wearing masks even while singing, increased natural or artificial ventilation, or moving activities outdoors.
Figure:
Note: Adapted from Katelaris et al. Diagram of church layout showing seating locations of index and secondary cases during an outbreak of SARS-CoV-2. Primary case is indicated in Red. Cases indicated in Green and Dark Blue confirmed by video recordings. Cases in Light Blue described their location via interview. Open access journal; all content freely available.
S-gene target failure as a marker of variant B.1.1.7 among SARS-CoV-2 isolates in the greater Toronto area, December 2020 to March 2021external icon. Brown et al. JAMA (April 8, 2021).
Key findings:
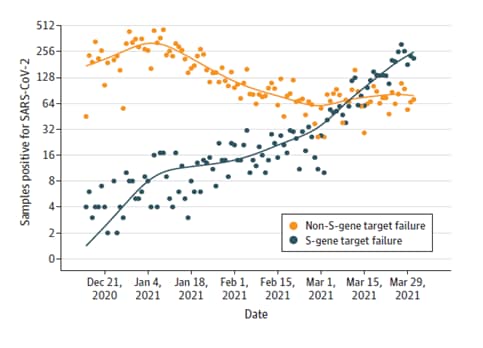
- The B.1.1.7 variant grew from 2% to 75% of daily cases as estimated by the surrogate measurement of S-gene target failure (SGTF).
- SGTF cases increased from 176 to 254 per day during the week of March 25-31 (weekly growth rate, 1.45; 95% CI, 1.18-1.75), while non-SGTF cases decreased from 81 to 79 per day (weekly growth rate, 0.97; 95% CI, 0.80-1.18) (Figure).
Methods: In Toronto, Canada, 20,051 positive NP swabs were taken between December 15, 2020 and March 31, 2021; 4,692 swabs were SGTF, i.e. positive for the N-gene and ORF1ab gene but negative for the S-gene. Confirmatory testing for B.1.1.7 in SGTF samples was done on a subset. Limitations: SGTF as a measurement of B.1.1.7 prevalence may not be accurate in the presence of other circulating variants with deletions in the S gene.
Implications: The B.1.1.7 variant has outpaced other variants in Toronto and can be tracked efficiently by SGTF prevalence.
Figure:
Note: Adapted from Brown et al. Daily Counts of SGTF (n = 4,692) vs non-SGTF (n = 15,359) samples among all SARS-CoV-2 clinical samples received between December 15, 2020, and March 31, 2021. Reproduced with permission from JAMA, 2021. Published online April 8, 2021. doi:10.1001/jama.2021.5607. Copyright© 2021 American Medical Association. All rights reserved.
Variants of Concern (VOCs) include SARS CoV-2 strains that have demonstrated increased transmissibility, more severe disease (hospitalization and death), or reduced therapeutic effectiveness or neutralization (in convalescent or vaccinee sera). The following studies examined the rate of household transmission for VOCs that have recently become more prevalent in US communities.
PEER-REVIEWED
A. Estimation of secondary household attack rates for L452R SARS CoV-2 variants detected by genomic surveillance at a community-based testing site in San Franciscoexternal icon. Peng et al. Clinical Infectious Diseases (March 31, 2021).
Key Findings:
- Household contacts exposed to West Coast variants (B.1.427 and B.1.429) were at higher risk of infection compared to household contacts exposed to lineages lacking these variants (0.36 vs 0.29, aRR = 1.25; 95% CI:0.98-1.59).
- Viral loads were similar among persons infected with West Coast (mean Ct 23.56; IQR 6.4) versus non-West Coast (mean Ct 23.67; IQR 7.8) variants, as was the proportion of individuals with symptoms (60.9% vs 64.3%).
Methods: SARS-CoV-2 genomic sequences were generated (n = 928) from positive samples (N = 1099) collected at a walk-up, rapid testing site during November 22-December 1, 2020 and January 10-29, 2021. Secondary household attack rates and mean sample viral loads were compared across West Coast and non-West Coast variants for the January samples only. Limitations: Potential misclassification of secondary cases as index cases; attack rate calculations did not account for other potential sources of infection; specimens collected before proliferation of other VOCs more recently identified in the US.
PREPRINTS (NOT PEER-REVIEWED)
B .Increased household secondary attack rates with variant of concern SARS-CoV-2 index casesexternal icon. Buchan et al. medRxiv (April 5, 2021). Published in Clinical Infectious Diseases as Increased household secondary attacks rates with variant of concern Severe Acute Respiratory Syndrome Coronavirus 2 index casesexternal icon (June 9, 2021).
Key findings:
- Across demographic categories, the secondary attack rate for variant of concern (VOC) (25.9%) index cases was consistently higher than for non-VOC (20.5%, p<0.01) index cases.
- In a matched cohort, the secondary attack rate for VOC index cases was 1.31 times higher than non-VOC index cases (aRR = 1.31, 95%CI 1.14-1.49).
- The higher secondary attack rate for VOC compared to non-VOC index cases was observed for asymptomatic (aRR = 1.91, 95% CI 0.96-3.80) and presymptomatic (aRR = 3.41, 95%CI 1.13-10.26) cases.
Methods: Index cases with VOC (n = 1318) reported from February 7 – 27, 2021 were 1:1 matched to non-VOC index cases. VOC index cases were confirmed for B.1.1.7 using whole genome sequencing or screened positive for the N501Y mutation using real-time PCR. Secondary household attack rates were the number of household cases that occurred 1-14 days after the index case divided by the number of household contacts. Limitations: Potential misclassification of secondary cases as index cases; small samples in some subgroups.
Implications for both studies (Buchan et al. and Peng et al.): Higher secondary attack rates related to VOCs contribute to increased cases of COVID-19. Increased transmissibility is not necessarily associated with viral load or presence of symptoms. Enhanced genomic surveillance paired with other mitigation and response capacity are important to identify and understand the clinical implications of new variants.
Detection, Burden, and Impact
- Masiá et al. Incidence of delayed asymptomatic COVID-19 recurrences in a 6-month longitudinal studyexternal icon. Journal of Infection (March 29, 2021). Among 5 COVID-19 patients who tested RT-PCR positive >90 days since first diagnosis, asymptomatic recurrent positivity occurred but did not necessarily represent a new infection.
- Kidman et al. Estimates and projections of COVID-19 and parental death in the USexternal icon. JAMA Pediatrics (April 5, 2021). More than 37,300 children in the US have lost at least 1 parent due to COVID-19, disproportionately affecting non-Hispanic Black children; bereavement support will be needed for these affected populations.
- Preston et al. Characteristics and disease severity of US children and adolescents diagnosed with COVID-19external icon. JAMA Network Open (April 9, 2021). Pediatric patients (N = 756) who were hospitalized with severe COVID-19 between March 1 and October 31, 2020, were more likely to have ≥1 chronic condition, to be aged 2-11 years, and male compared to hospitalized pediatric patients without severe COVID-19; there was no association between race/ethnicity or insurance type and severe COVID-19.
Transmission of SARS-CoV-2
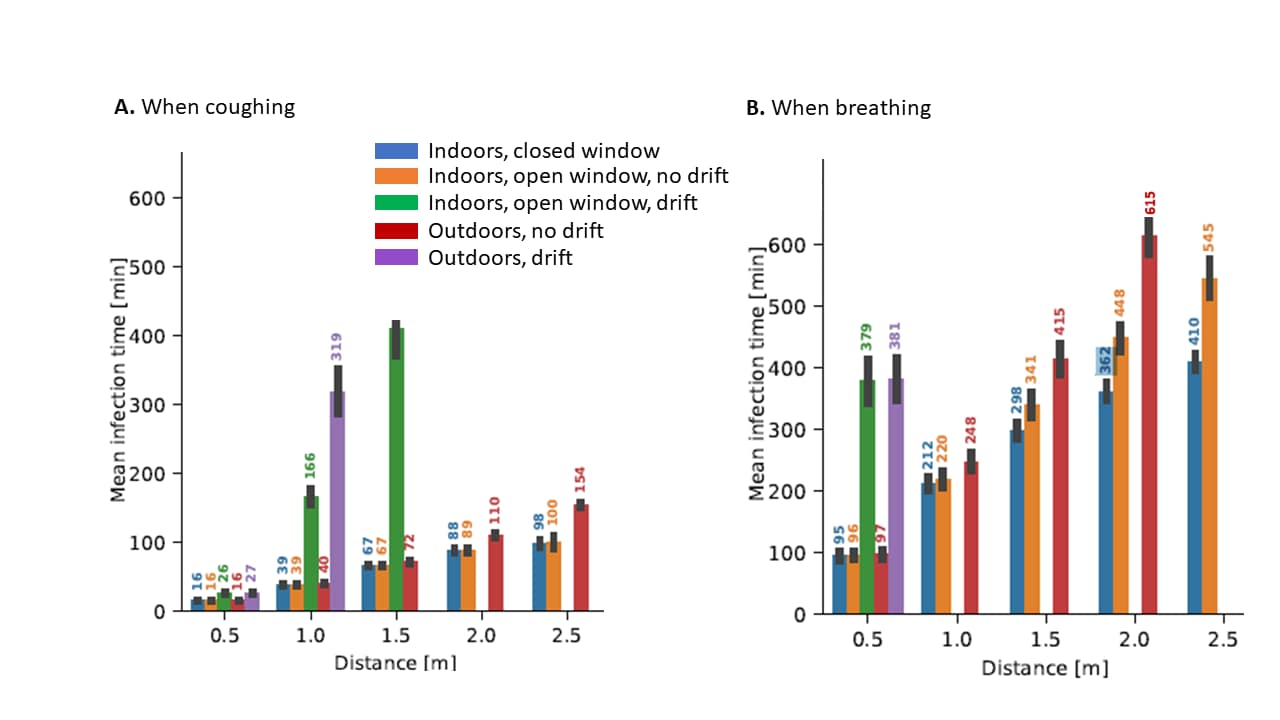
- Dobramysl et al. Mean time to infection by small diffusing droplets containing SARS-CoV-2 during close social contactsexternal icon. medRxiv (Preprint, April 7, 2021). Beyond factors such as exposure time, physical proximity, air flow dynamics, and mask use, time to infection via aerosol particles may be strongly affected by whether the infected person is coughing.
Figure:
Note: Adapted from Dobramysl et al. Mean infection times for various scenarios when an infected person is coughing once every 5 minutes [Panel A] and an infected person is breathing regularly [Panel B]. Missing bars indicate scenarios in which estimated infection times extended beyond 12 hours. Used by permission of authors.
Natural History of SARS-CoV-2 Infection
- Fournier et al. Emergence and outcome of the SARS-CoV-2 “Marseille-4” variantexternal icon. International Journal of Infectious Diseases (March 27, 2021). Genomic analysis of the Marseille-4 variant identified 13 hallmark mutations appearing as a set; among 2,106 diagnoses in September 2020, the variant was associated with greater hypoxemia than clade 20A strains and caused several cases of re-infection. A nearby mink farm was suggested as possible source.
- Nuñez et al. SARS-CoV-2 B.1.1.7 infection of Syrian hamster does not cause more severe disease and is protected by naturally acquired immunityexternal icon. bioRxiv (Preprint, April 2, 2021). Published in mSphere as SARS-CoV-2 B.1.1.7 infection of Syrian hamster does not cause more severe disease, and naturally acquired immunity confers protectionexternal icon (June 16, 2021). In hamsters, the B.1.1.7 variant did not differ from the B.1 variant in pathogenicity, and infection from the original D614 variant conferred protection against a secondary challenge with B.1.1.7.
- Fischer et al. ChAdOx1 nCoV-19 (AZD1222) protects hamsters against SARS-CoV-2 B.1.351 and B.1.1.7 diseaseexternal icon. bioRxiv (Preprint, March 15, 2021). Published in Nature Communicationsexternal icon (October 7, 2021). Compared to controls, vaccinated hamsters showed less weight loss, fewer lung lesions, and no infectious virus in the lungs or pulmonary pathology, providing evidence that this vaccine reduces clinical disease due to B.1.1.7 and B.1.351 variants in hamsters.
Prevention, Mitigation, and Intervention Strategies
- Beharier et al. Efficient maternal to neonatal transfer of SARS-CoV-2 and BNT162b2 antibodiesexternal icon. medRxiv (Preprint, April 6, 2021). Published in the Journal of Clinical Investigation as Efficient maternal to neonatal transfer of antibodies against SARS-CoV-2 and BNT162b2 mRNA COVID-19 vaccineexternal icon (May 20, 2021). The BNT162b2 vaccine elicited strong maternal humoral response that reached similar levels in the fetus within 15 days following the first dose and were similar to levels occurring with sensitization (vaccine vs. infection), providing evidence to support vaccination during pregnancy.
- Sattler et al. Impaired humoral and cellular immunity after SARS-CoV2 BNT162b2 (Tozinameran) prime-boost vaccination in kidney transplant recipientsexternal icon. medRxiv (Preprint, April 7, 2021). Published in the Journal of Clinical Investigationexternal icon (June 8, 2021). Humoral and cellular responses to vaccination were much lower in kidney transplant recipients compared with healthy vaccinees and hemodialysis patients, suggesting a possible need for revised vaccination approaches in immunosuppressed patients.
- Krogstad et al. No evidence of infectious SARS-CoV-2 in human milk: analysis of a cohort of 110 lactating womenexternal icon. medRxiv (Preprint, April 7, 2021). Published in Pediatric Research as No infectious SARS-CoV-2 in breast milk from a cohort of 110 lactating womenexternal icon (January 19, 2022). The breast milk of 110 women recently infected with COVID-19 infrequently contained SARS-CoV-2 RNA (6%), with no evidence of infectious virus, suggesting that breastfeeding is unlikely to cause infection in infants.
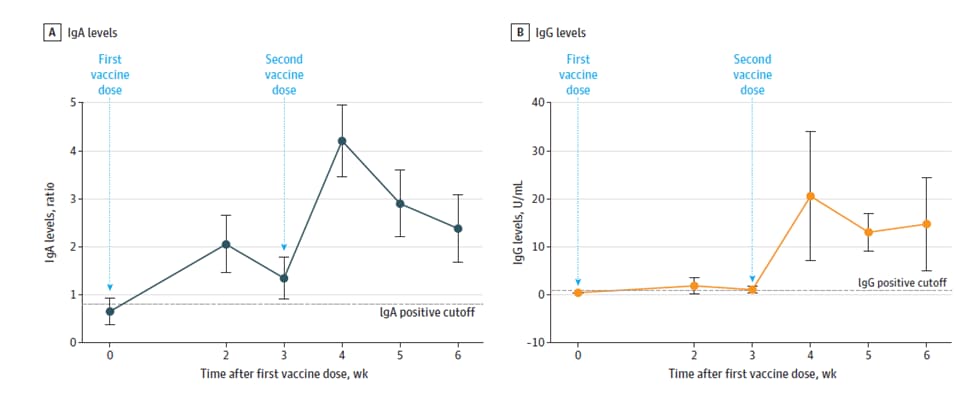
- Perl et al. SARS-CoV-2–specific antibodies in breast milk after COVID-19 vaccination of breastfeeding womenexternal icon. JAMA (April 12, 2021). SARS-CoV-2 specific IgA and IgG antibodies were found in breast milk from 84 women 6 weeks after Pfizer-BioNTech vaccination. No serious adverse events were found in infants or women post vaccination.
Figure:
Note: Adapted from Perl et al. Change in IgA and IgG levels in breast milk over time. Mean and 95% CIs shown. Reproduced with permission from JAMA, 2021. Published online April128, 2021. doi:10.1001/jama.2021.5782. Copyright© 2021 American Medical Association. All rights reserved.
Disclaimer: The purpose of the CDC COVID-19 Science Update is to share public health articles with public health agencies and departments for informational and educational purposes. Materials listed in this Science Update are selected to provide awareness of relevant public health literature. A material’s inclusion and the material itself provided here in full or in part, does not necessarily represent the views of the U.S. Department of Health and Human Services or the CDC, nor does it necessarily imply endorsement of methods or findings. While much of the COVID-19 literature is open access or otherwise freely available, it is the responsibility of the third-party user to determine whether any intellectual property rights govern the use of materials in this Science Update prior to use or distribution. Findings are based on research available at the time of this publication and may be subject to change.