COVID-19 Science Update released: February 19, 2021 Edition 77

The COVID-19 Science Update summarizes new and emerging scientific data for public health professionals to meet the challenges of this fast-moving pandemic. Weekly, staff from the CDC COVID-19 Response and the CDC Library systematically review literature in the WHO COVID-19 databaseexternal icon, and select publications and preprints for public health priority topics in the CDC Science Agenda for COVID-19 and CDC COVID-19 Response Health Equity Strategy.
Section headings in the COVID-19 Science Update have been changed to align
with the CDC Science Agenda for COVID-19.
Here you can find all previous COVID-19 Science Updates.
PEER-REVIEWED
Rapid decline of neutralizing antibodies against SARS-CoV-2 among infected healthcare workers.external icon Marot et al, Nature Communications, (February 8, 2021).
Key findings:
- Healthcare workers (HCWs) infected with mild symptomatic SARS-CoV-2 infection experienced changes in IgA profiles that were associated with declining neutralizing activity 2 months after symptom onset.
- Anti-receptor binding domain (RBD) IgA decreased from 100% at baseline to 38.5% at 3 months, but anti-RBD immunoglobulin G (IgG) remained relatively stable (100% at baseline to 92.3% at 3 months).
- Significant concomitant decreases in neutralizing antibody titers (NAbs) were observed between day 21 and 2 months and 3 months, respectively, suggesting that NAbs correlate with IgA.
Methods: Observational study of serum samples of 26 HCW recovering from mild, RT-PCR confirmed SARS-CoV2 infection to assess the development of protective antibody response and duration. Changes in levels of IgG, IgA, and IgM antibodies and NAbs assessed at 3 time points post-symptom onset: day 21, 2 months, and 3 months. Limitations: Small sample size.
Implications: Short-lasting immune protection suggests prevention and control measures should be maintained among seropositive HCWs.
PEER-REVIEWED
Emergence of a novel SARS-CoV-2 variant in Southern Californiaexternal icon. Zhang et al. JAMA (February 11, 2021).
Key findings:
- In California, diverse lineages with 2 main clusters, the small 20G cluster (22%) and a larger, novel SARS-CoV-2 variant (CAL.20C) cluster (36%), have been identified.
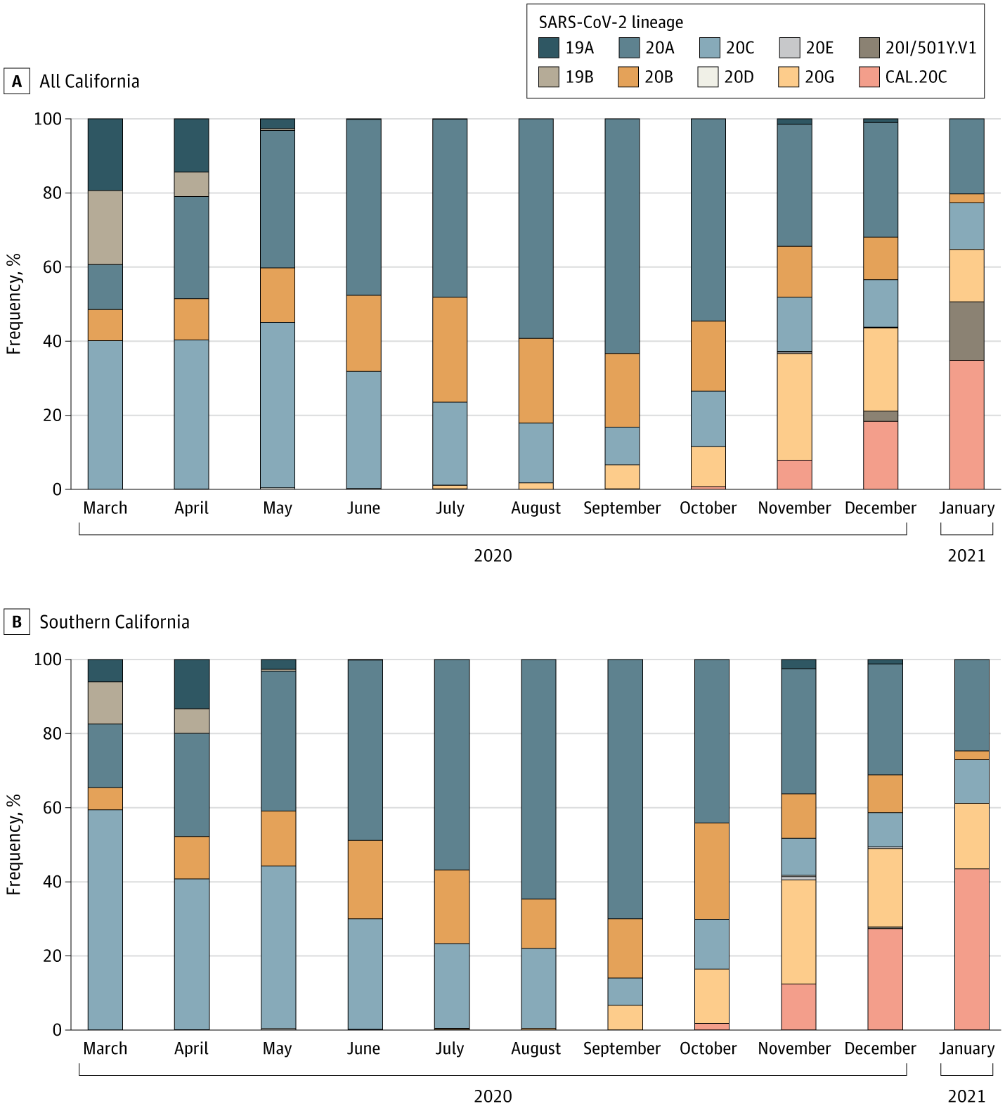
- 20C was first identified in 1 sample from Los Angeles County in July 2020 and accounted for 35% of all California and 44% of all Southern California strains in January 2021 (Figure).
Methods: Of 2,311 SARS-CoV-2 specimens with Ct <30 from symptomatic hospitalized and ambulatory care patients at Cedars-Sinai Medical Center in Los Angeles from November 22−December 28, 2020, 185 randomly selected specimens underwent sequencing and Nexstrain phylogenetic analysis along with 1,480 globally representative genomes present in GISAID as of January 11, 2021. Prevalence of clades in 10,431 samples from California, including 4,829 from Southern California, from March 4, 2020 to January 22, 2021 in GISAID were described. Limitations: Selection bias as analyses were limited to publicly available genomes and a comparatively small set of local samples.
Implications: A novel SARS-CoV-2 variant (CAL.20C) emerged in Southern California in 2020 and has increased in prevalence. Studies are warranted to assess the impact of this variant on disease outcomes and transmission.
Figure.
Note: from Zhang et al. Timeline of variants from March 2020 to January 2021 among all SARS-CoV-2 samples in California (A) and Southern California (B). The CAL.20C variant is in orange. Reproduced with permission from JAMA, Emergence of a novel SARS-CoV-2 variant in Southern California, 2021. doi:10.1001/jama.2021.1612. Copyright© (2021) American Medical Association. All rights reserved.
PREPRINTS (NOT PEER-REVIEWED)
Detection of genomic variants of SARS-CoV-2 circulating in wastewater by high-throughput sequencing. external iconPérez-Cataluña et al. medRxiv. (February 10, 2021).
Key findings:
- A total of 238 nucleotide substitutions and 6 deletions were found in SARS-CoV-2 isolates.
- 3 novel variants with spike gene mutations (G648V, A893T, and L1152S) were identified.
- 6 variants with known spike gene mutations were detected for the first time in Spain.
Methods: 40 samples were collected from April to October 2020 at 14 wastewater treatment plants located in 3 geographical regions within the Spanish peninsula for a genomic sequencing study. Variant analysis was conducted to detect genetic differences between circulating strains and the reference genome, SARS-CoV-2 isolate Wuhan-Hu-I. Limitations: Detection of variants in wastewater depends on population prevalence of isolate; small sample size.
Implications: Wastewater sequencing facilitates real-time detection of emerging variants of SARS-CoV-2 that might be missed by sequencing of samples from individuals seen in clinical settings. These findings are relevant for surveillance and monitoring efforts.
PEER-REVIEWED
Measuring the impact of COVID-19 vaccine misinformation on vaccine intent in the UK and USA.external icon Loomba et al. Nature Human Behaviour (February 5, 2021).
Key findings:
- Intent to vaccinate for one’s own protection dropped after exposure to misinformation from 42.5% (95% percentile intervals [PI]: 41.0%−44.1%) to 39.8% (PI: 38.2%−41.5%) in the USA.
- Intent to vaccinate was higher if the rationale was to protect others but showed similar declines after exposure to misinformation from 53.3% (PI: 51.8%−54.9%) to 46.4% (PI: 44.8%−48.1%) in the USA.
Methods: Randomized, controlled trial of 8,001 participants recruited from an online panel with quotas based on national demographic patterns for gender, age, and sub-national region, from September 7-14, 2020. The treatment group (n = 6,001) was exposed to COVID-19 vaccine misinformation based on recently circulating online imagery; the control group (n = 2,000) was exposed to factual information about the vaccine. Participants reported intention to vaccinate if a vaccine were available before and after exposure to online imagery. Limitations: Presentation of information and misinformation in the experimental setting might not represent exposure in real-world circumstances.
Implications: Exposure to misinformation lowers intent to vaccinate both for individual protection and protection of others. Health communications efforts must counter the effects of misinformation as well as present the benefits of vaccination against COVID-19.
PEER-REVIEWED
Genomic Signatures of SARS-CoV-2 Associated with Patient Mortalityexternal icon. Dumonteil et al. Viruses (February 2, 2021).
Key findings:
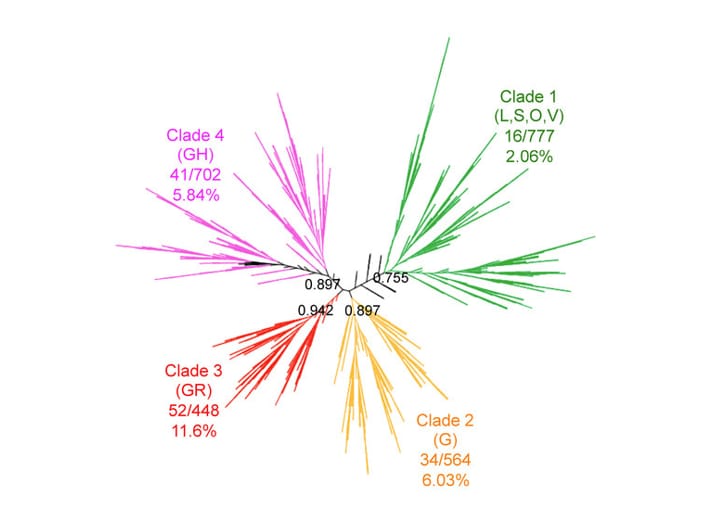
- Phylogenetic analysis of SARS-CoV-2 genome sequences revealed 4 well-resolved clades.
- Overall mortality was 5.7% and differed significantly between the clades ranging from 2.06% for Clade 1 to 11.6% for Clade 3 (Figure).
- 3 previously undetected single-nucleotide polymorphisms (SNPs) in the SARS-CoV-2 genome were associated with patient mortality after adjusting for patient age, sex, geographic region, and month.
Methods: Phylogenetic analysis of 2,508 whole-genome sequences from SARS-CoV-2 virus selected from the GISAIDexternal icon database from December 2019−June 2020 and linked with metadata including disease severity and demographics. A phylogenetic tree was constructed from SNPs identified using Geneious 11. Generalized linear models were used to test for associations between mortality rates, viral clades, and co-variates. Limitations: Sequenced viral genomes are not a random sample of the global virus population and thus subject to selection bias; comorbidities were not included in the model.
Implications: The SARS-CoV-2 SNPs associated with mortality warrant further investigation to understand mechanisms underlying COVID-19 morbidity and mortality.
Figure.
Note. Adapted from Dumonteil et al. Genomic diversity of SARS-CoV-2 and patient mortality. Unrooted phylogenetic tree of SARS-CoV-2 genomes showing 4 main Clades (1-4) with a strong phylogenetic support as specified by the numbers at the base of each major branch and associated mortality rates which varied significantly between Clades (X2=47.93, p<0.00001). GISAIDexternal icon clade names are indicated in parenthesis. Licensed under CC BY 4.0.
A SARS-CoV-2 cluster in an acute care hospital.external icon Klompas et al. Annals of Internal Medicine (February 9, 2021).
Key findings:
- Discontinuation of isolation in a symptomatic index case with 2 negative SARS-CoV-2 RT-PCR results 12 hours apart led to 52 secondary infections; the patient was not diagnosed with COVID-19 until day 11.
- 7/52 (13%) secondary infections were detected only on second or subsequent serial tests.
- 6 /14 (43%) patients and 13/38 (34%) employees were asymptomatic.
- Risk factors for transmission to employees included presence during nebulization, caring for patients with dyspnea or cough, lack of eye protection, at least 15 minutes of exposure to SARS-CoV-2 patients, and interactions with SARS-CoV-2– positive employees in clinical areas.
- There were at least 2 patient-to-staff transmissions detected via whole-genome sequencing that occurred despite staff wearing both masks and face shields and in the absence of aerosol-generating procedures.
Methods: Whole-genome sequencing to identify SARS-CoV-2 clusters in large acute care hospital. SARS-CoV-2-positive and negative employees were enrolled in a case–control study to compare clinical interactions, personal protective equipment use, breakroom and workroom practices, and exposures outside the hospital. Limitations: Difficult to isolate exact transmission routes due to interconnections between patients and employees; recall bias may have led to either underestimation or overestimation of adherence to infection control practices
Implications: SARS-CoV-2 clusters can occur in hospitals despite robust infection control policies and may be contributed to by the inherent insensitivity of testing at specific points in infection time course. Surgical mask and face shields may not provide adequate protection for close and sustained encounters with highly contagious patients, especially if they cannot wear a mask.
PREPRINTS (NOT PEER-REVIEWED)
Poor antigen-specific responses to the second BNT162b2 mRNA vaccine dose in SARS-CoV-2-experienced individuals.external icon Samanovic et al. medRxiv (February 9, 2021). Published in Science Translational Medicine as Robust immune responses are observed after one dose of BNT162b2 mRNA vaccine dose in SARS-CoV-2-experienced individualsexternal icon (December 7, 2022).
Key findings:
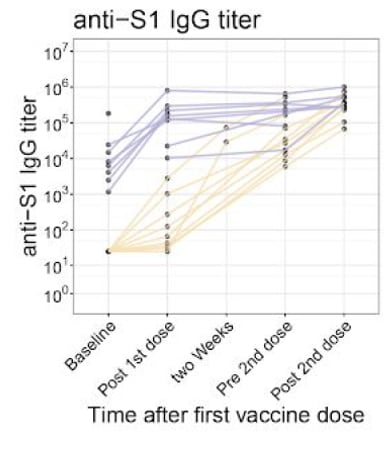
- After the first vaccine dose, median fold-change in S1 protein antibodies was significantly greater in SARS-CoV-2-experienced patients than SARS-CoV-2-naïve patients (47 vs 2.6) (Figure A).
- After the second vaccine dose, median fold-change was significantly lower in SARS-CoV-2-experienced patients than SARS-CoV-2-naïve patients (1.4 vs 13) (Figure A).
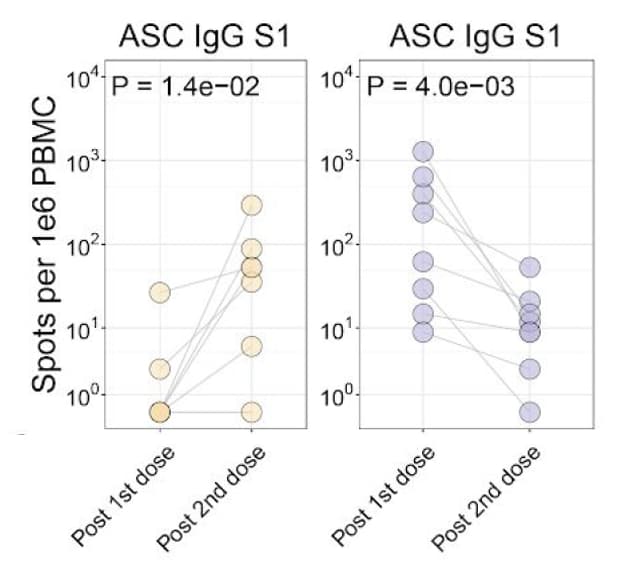
- Similarly, S1-protein antigen-specific antibody secreting cell (ASC) responses were robust in SARS-CoV-2-experienced patients after the first dose but minimal after the second dose and were minimal in SARS-CoV-2-naïve patients after the first dose but more robust after the second dose (Figure B).
Methods: 32 persons (19 SARS-CoV-2-naïve and 13 SARS-CoV-2-experienced) received two doses of BNT162b2 (Pfizer) mRNA vaccine. Samples were collected at various timepoints including baseline, after first dose, before second dose and after second dose. Antibody titers and ASC cell responses were quantified by ELISA and ELISpot, respectively. Limitations: Small sample size.
Implications: A single vaccine dose for persons with prior SARS-CoV-2 infection may offer sufficient protection. However, the extent of protection offered and whether T cell responses are adequately stimulated by one dose in these persons remains unclear. Another recent studyexternal icon observed similar findings after the first dose of the BNT162b2 (Pfizer) mRNA vaccine in healthcare workers with previous SARS-CoV-2 infection.
Figure:
Note: Adapted from Samanovic et al. (A) S1 antibody titers were assessed in SARS-CoV-2-naive and SARS-CoV-2 experienced subjects. (B) S1 direct enzyme-linked immunospot (ELISpot) assay results shown for SARS-CoV-2-naive or SARS-CoV-2 experienced subjects. Connected lines indicate repeated measurements of the same subjects. IgG = Immunoglobulin G; ASC = antibody secreting cell; PBMC = peripheral blood mononuclear cells. Licensed under CC-BY-NC-ND 4.0.
Efficacy of ChAdOx1 nCoV-19 (AZD1222) vaccine against SARS-CoV-2 VOC 202012/01 (B.1.1.7). external iconEmary et al. The Lancet (February 4, 2021). Published in The Lancet as Efficacy of ChAdOx1 nCoV-19 (AZD1222) vaccine against SARS-CoV-2 variant of concern 202012/01 (B.1.1.7): An exploratory analysis of a randomised controlled trialexternal icon (April 10, 2021).
Key findings:
- Vaccine efficacy against symptomatic COVID-19 caused by the B.1.1.7 variant was 74.6% (95% CI 41.6%-88.9%) compared with 84.1% (95% CI 70.7%-91.4%) against non-B.1.1.7 variant.
- Sera from ChAdOx1 nCoV-19 vaccinees showed a 9-fold reduction in neutralizing activity against the B.1.1.7 lineage compared with one non-B.1.1.7 variant.
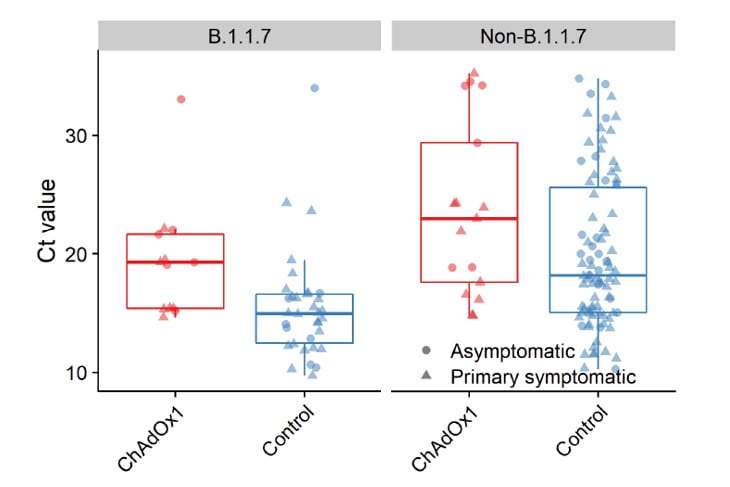
- Participants who received the ChAdOx1 nCoV-19 vaccine had significantly higher Ct values and reduced duration of RT-PCR-positivity (median reduction = 1 week) than those who received the control vaccine (Figure).
Methods: Ongoing, single-blind, multicentered randomized phase II/III vaccine trial for the ChAdOx1 nCoV-19 vaccine in adults ≥18 years of age in the United Kingdom. Participants were randomized to receive ChAdOx1 nCoV-19 vaccine or meningococcal vaccine (MenACWY). Vaccine efficacy analyses used samples from symptomatic (n=120) and asymptomatic/unknown (n=44) participants enrolled from October 1, 2020-January 14, 2021. Limitations: Laboratory testing was performed on self-administered nasal and throat swabs; not all samples were processed. Small sample size.
Implications: The ChAdOx1 nCoV-19 vaccine provides similar efficacy against symptomatic COVID-19 disease caused by the B.1.1.7 as non-B.1.1.7 variants but reduced neutralization activity. Reduced viral load and viral shedding was seen among persons vaccinated with ChAdOx1 nCoV-19, this could lead to reduced disease transmission. These data support ongoing implementation of mass vaccination.
Figure:
Note: Adapted from Emary et al. Minimum Ct values across all PCR positive swabs. Ct values from positive PCR tests performed at Lighthouse laboratories using a ThermoFisher TaqPath 3-gene assay. Each data point represents one participant. Low PCR Ct values are associated with higher viral load. P-values from Wilcoxon Rank Sum tests comparing ChAdOx1 nCoV-19 with Control are: B.1.1.7 variant p = 0.0113, non-B.1.1.7 variants p = 0.0201. Used by permission of publisher.
PREPRINTS (NOT PEER-REVIEWED)
Increasing prevalence of SARS-CoV-2 variant strains has raised concerns about both the efficacy of currently available COVID-19 vaccines and reinfection among individuals who have recovered from SARS-CoV-2 infection. Two studies report on antibody responses to SARS-CoV-2 variants among healthy individuals who received the vaccine (Tada et al.external icon), recovered individuals who have not received the vaccine (Tada et al.external icon), and recovered individuals who have received one dose of vaccine (Stamatatos et al.external icon).
A. Neutralization of viruses with European, South African, and United States SARS-CoV-2 variant spike proteins by convalescent sera and BNT162b2 mRNA vaccine-elicited antibodies.external icon Tada et al. bioRxiv. (February 7, 2021). Published in mBio as Convalescent-phase sera and vaccine-elicited antibodies largely maintain neutralizing titer against global SARS-CoV-2 variant spikesexternal icon (June 1, 2021).
Key findings:
- Vaccination is effective in protecting against B.1.1.7 but there was a 3-fold reduction in neutralizing titers for B.1.351, likely due to the E484K mutation.
- For D614G, neutralizing titers among vaccinated individuals were 7-fold greater compared with titers among recovered individuals.
- Convalescent sera neutralized several variant viruses, including B.1.1.7 and B.1.351, with titers similar to those measured against D614G.
Methods: Blood and isolated serum collected from 10 individuals recovered from SARS-CoV-2 and 5 healthy individuals vaccinated with the BNT162b2 (Pfizer-BioNTech) mRNA vaccine (days 0, 7, and 28 days post-vaccination) were used to measure antibody neutralization titers against variants including D614G, B.1.1.7, B.1.351, COH.20G/677H Columbus Ohio, 20A.EU2 Europe, and mink cluster 5. Limitations: Small sample size.
B. Antibodies elicited by SARS-CoV-2 infection and boosted by vaccination neutralize an emerging variant and SARS-CoV-1.external icon Stamatatos et al. medRxiv. (February 8, 2021). Published in Science as mRNA vaccination boosts cross-variant neutralizing antibodies elicited by SARS-CoV-2 infectionexternal icon (March 25, 2021).
Key findings:
- Convalescent sera displayed neutralizing activity against both the Wuhan-Hu-1 and B.1.351 strains, but less against B.1.351, 4-8 months post-SARS-CoV-2 infection.
- Following a single dose of either the Pfizer-BioNTech or Moderna mRNA vaccine, neutralizing antibody (NAbs) titers were boosted approximately up to 1000-fold for each strain.
- NAbs titers were 2–3-fold lower for B.1.351 compared to Wuhan-Hu-1.
Methods: Blood and isolated serum were collected from 10 recovered SARS-CoV-2 persons. Antibody binding and neutralization titers were measured before (mean = 202 days post-symptom onset) and after (mean = 16 days post-vaccination) receiving a single dose of either the BNT162b2 (Pfizer-BioNTech) or mRNA-1273 (Moderna) vaccine. Limitations: Small sample size.
Implications for both studies (Tada et al. and Stamatatos et al.): Current COVID-19 mRNA vaccines are effective in protecting against variants among persons previously infected and uninfected with SARS-CoV-2. However, mRNA vaccines likely offer reduced protection against B.1351 in some individuals. Although persons previously infected by SARS-CoV-2 have some protection against variants, they might benefit from boosted vaccination.
Detection, Burden, and Impact
- Bruckner et al. Estimated seroprevalence of SARS-CoV-2 antibodies among adults in Orange County, California.external icon Scientific Reports (February 4, 2021). Active surveillance of 2,979 persons who completed a survey and were later tested for SARS-CoV-2 at drive-thru sites between July 10 and August 16, 2020, found a seroprevalence of 11.5%; these results imply that SARS-CoV-2 infections might be underreported.
- Geriatric Medicine Research Collaborative. Age and frailty are independently associated with increased COVID-19 mortality and increased care needs in survivors: results of an international multi-centre study.external icon Age and Ageing (February 5, 2021). Analysis of data from 5,711 hospitalized adults in 12 countries in a multi-center cohort study found that increasing age (hazard ratio [HR] 3.57, CI 2.54-5.02) and severity of frailty (HR 3.03, CI 2.29-4.00) were independently associated with increased risk of death from COVID-19.
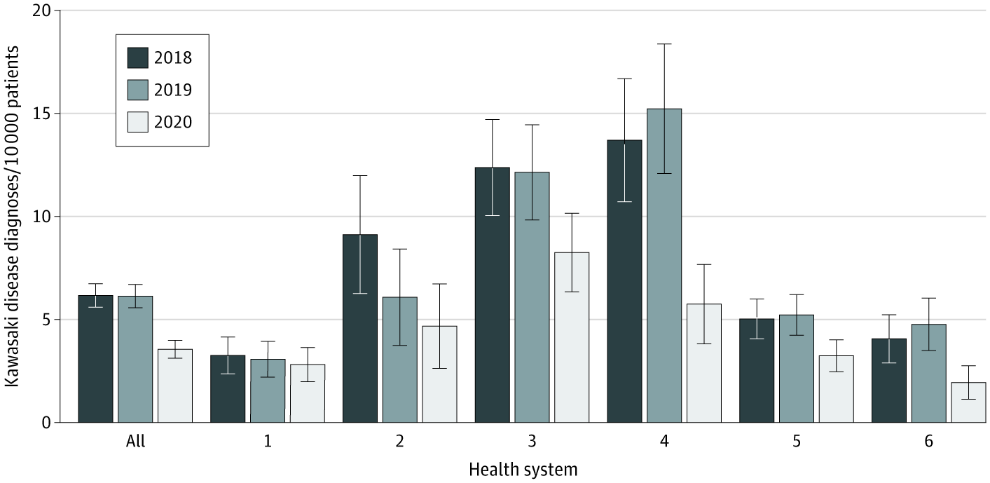
- Bailey et al. Assessment of 135,794 pediatric patients tested for severe acute respiratory syndrome coronavirus 2 across the United States.external icon JAMA Pediatrics (November 23, 2020). In this retrospective cohort study of persons <25 years old (PEDSnet), increasing age, chronic conditions, and non-White race/ethnicity were associated with higher rates of SARS-CoV-2 infection; this study also noted a 40% decline in rates of Kawasaki Disease (KD) from March 1-May 15 2020 compared with same intervals in 2018 and 2019, suggesting that KD may not be a good proxy for multisystem inflammatory syndrome.
Figure.
Note: Adapted from Bailey et al. Rates of Kawasaki Disease Diagnosis in the PEDSnet Population. Between March 1 and May 15 in 2018 and in 2019, the mean number of patients seen was used to establish an at-risk denominator and case counts estimated based on diagnoses assigned during this date interval were taken from PEDSnet data for 2018, 2019, 2020. Vertical bars indicate 95% CIs. Reproduced with permission from JAMA Pediatrics, Assessment of 135,794 pediatric patients tested for severe acute respiratory syndrome coronavirus 2 across the United States. 2021. doi:10.1001/jamapediatrics.2020.5052. Copyright© (2021) American Medical Association. All rights reserved.
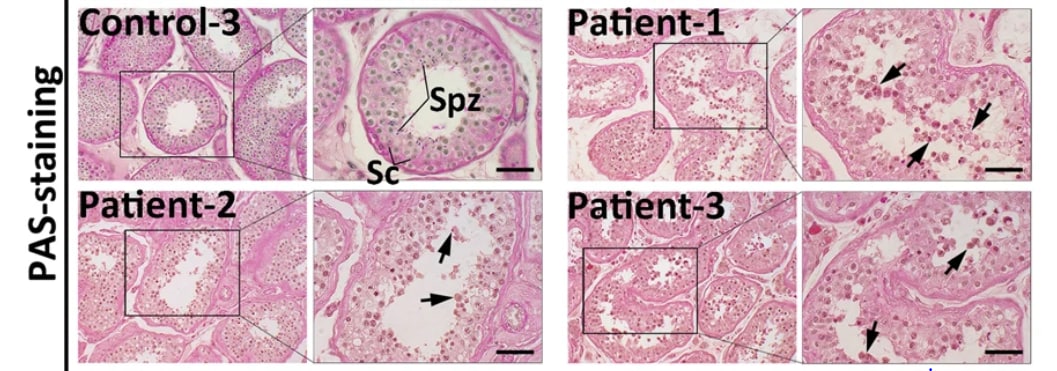
- Ma et al. Pathological and molecular examinations of postmortem testis biopsies reveal SARS-CoV-2 infection in the testis and spermatogenesis damage in COVID-19 patients.external icon Cellular & Molecular Immunology (December 14, 2020). Examination of testicular biopsy samples from 5 COVID-19 cases showed significant degeneration of germ cells and upregulation of 32 inflammatory cytokines compared with 3 uninfected controls, illustrating the potential impact of SARS-CoV-2 infection on male fertility.
Figure.
Note: Adapted from Ma et al. Spermatogenesis damage was observed in COVID-19 patients. Histological analyses using periodic acid-Schiff (PAS) staining of testicular sections from COVID-19 patients (patients 1, 2, and 3) shows numerous degenerated germ cells (arrows) sloughing into the lumen of the seminiferous tubules; normal spermatogenesis was observed in control patients (control 3). Spz = spermatozoa, Sc = Sertoli cells. Scale bar = 100 μm. Permission request in process.
Transmission of SARS-CoV-2
- Tarhini et al. Long term SARS-CoV-2 infectiousness among three immunocompromised patients: from prolonged viral shedding to SARS-CoV-2 superinfeexternal icon The Journal of Infectious Diseases (February 8, 2021). Case series of 3 severely immunocompromised COVID-19 patients who shed infectious virus for up to 4 months after symptom onset suggests the need for assessing infectiousness with viral culture and/or Ct values prior to discontinuing isolation in this population.
- Levine-Tiefenbrun et al. Decreased SARS-CoV-2 viral load following vaccination.external icon medRxiv (Preprint, February 8, 2021). Published in Nature Medicine as Initial report of decreased SARS-CoV-2 viral load after inoculation with the BNT162b2 vaccineexternal icon (March 29, 2021). By examining Ct values over time after the first dose of the BNT162b2 (Pfizer-BioNTech) mRNA vaccine among 2,897 persons with post-vaccination SARS-CoV-2 infection, the authors found a 4-fold reduction in mean viral load on days 12–28 post vaccination; these data suggest that the vaccine reduces infectiousness and thus limits further SARS-CoV-2 transmission.
Social, Behavioral, and Communication Science
- Robertson et al. Did people’s behavior after receiving negative COVID-19 tests contribute to the spread?external icon Journal of Public Health (February 8, 2021). Analysis of COVID-19 hospitalization and testing data from 25 countries throughout 2020 suggests that persons who tested falsely negative assumed they were at reduced risk and engaged in activities that increased the spread of SARS-CoV-2.
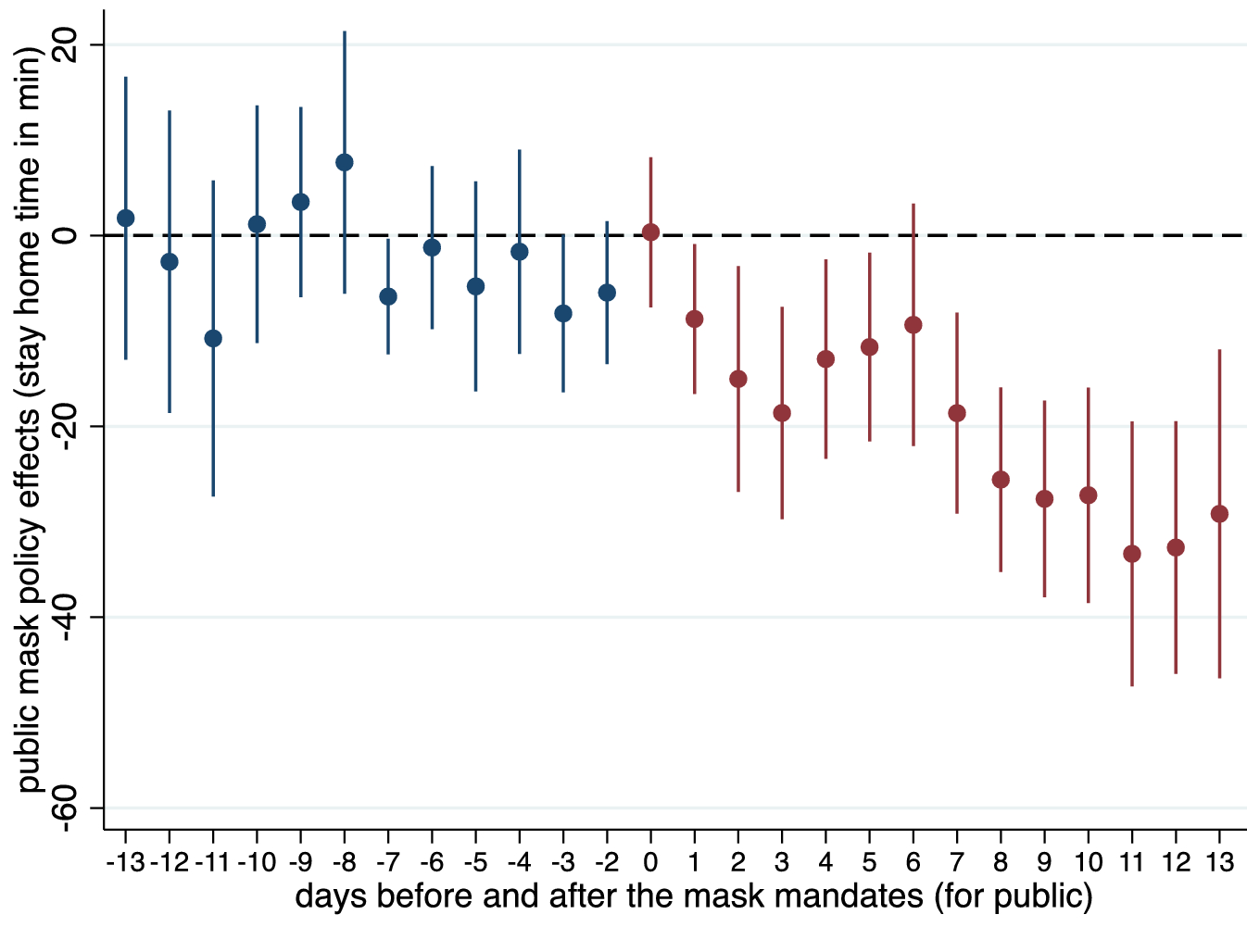
- Yan et al. Risk compensation and face mask mandates during the COVID-19 pandemic.external icon Scientific Reports (February 4, 2021). Using SafeGraph smart device location and mask mandate data, the authors found that face mask orders led to less time spent at home and more visits to risky locations such as restaurants which is beneficial for the economy but might increase transmission of SARS-CoV-2.
Figure.
Note: Adapted from Yan et al. Change in time spent at home, in minutes, 14 days before and after public mask mandate implementation (day 0). Licensed under CC BY 4.0.
Natural History of SARS-CoV-2 Infection
- Brookman et al. Effect of the new SARS-CoV-2 variant B.1.1.7 on children and young people.external icon Lancet Child and Adolescent Health (February 10, 2021). Data from the first (March 1–May 31, 2020) and second (November 1, 2020–January 19, 2021) waves of the pandemic at one hospital in south London suggest that SARS-CoV-2 infection with the B.1.1.7 variant does not result in a more severe disease course than the wild-type SARS-CoV-2 strain and that severe COVID-19 remains uncommon in children and young people.
Prevention, Mitigation, and Intervention Strategies
- Shimabukuro et al. Reports of anaphylaxis after receipt of mRNA COVID-19 vaccines in the US—December 14, 2020–January 18, 2021.external icon JAMA Insights (February 12, 2021). From December 14, 2020–January 18, 2021, 66 reports of anaphylaxis were identified that met the Brighton Collaboration case definition in the Vaccine Adverse Event Reporting Systemexternal icon: 47 followed the BNT162b2 (Pfizer-BioNTech) mRNA vaccine for a rate of 4.7 cases/million doses administered and 19 followed the mRNA-1273 (Moderna) vaccine for a rate of 2.5 cases/million doses administered; these data confirm that post-vaccination anaphylaxis is a rare event.
Disclaimer: The purpose of the CDC COVID-19 Science Update is to share public health articles with public health agencies and departments for informational and educational purposes. Materials listed in this Science Update are selected to provide awareness of relevant public health literature. A material’s inclusion and the material itself provided here in full or in part, does not necessarily represent the views of the U.S. Department of Health and Human Services or the CDC, nor does it necessarily imply endorsement of methods or findings. While much of the COVID-19 literature is open access or otherwise freely available, it is the responsibility of the third-party user to determine whether any intellectual property rights govern the use of materials in this Science Update prior to use or distribution. Findings are based on research available at the time of this publication and may be subject to change.